HiC-TAD Library Media Gallery
Chromatin 3D Structure Visualizations & Hi-C Analysis Results
Lab Presentation Slides
Documentation NEW
Comprehensive documentation walkthroughs and analysis guides.
Full Analysis Reports
Standalone HTML reports with all figures, tables, and biological interpretation. Open in a new tab for the complete analysis.
Synthesis Experiments NEW
In-silico chromatin engineering
Unc5b · CTCF deletions · Synthetic insulator
Drug Discovery Pipeline NEW
Genomics → Protein → Structure → Molecules
UNC5B · ESM-2 · ESMFold · MolMIM · DiffDock
Dovetail® Analysis Suite NEW
Full pipeline: Loops · Compartments
HiChIP · Capture · SVs · CNVs · Phasing
LinkPrep™ Analysis
Dovetail Tn5 proximity ligation
TADs · SVs · CNVs · Phasing
AlphaGenome Analysis
Full insulator deletion report
Both Edward & Jingyun — all cell types
Deletion Scan — Edward chr12
12-site sensitivity scan
Original insulator size (~3 kb)
Deletion Scan — Jingyun chr13
12-site sensitivity scan
Original insulator size (~5.3 kb)
Multi-Size Scan — Edward chr12
10, 40, 80 kb deletions
12 positions × 3 sizes
Multi-Size Scan — Jingyun chr13
10, 40, 80 kb deletions
12 positions × 3 sizes
Interactive 3D Visualizations NEW
Static PNG Visualizations
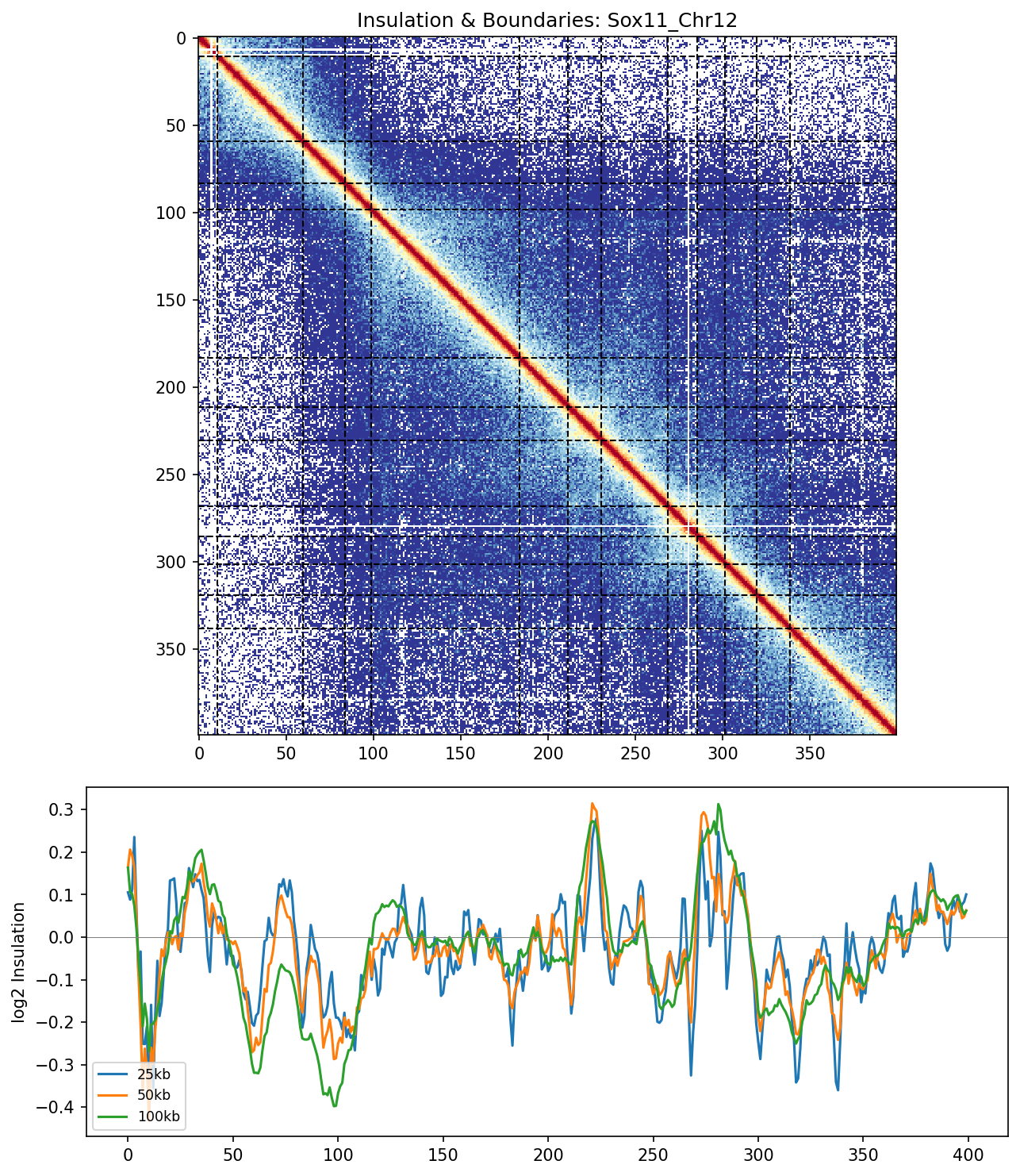
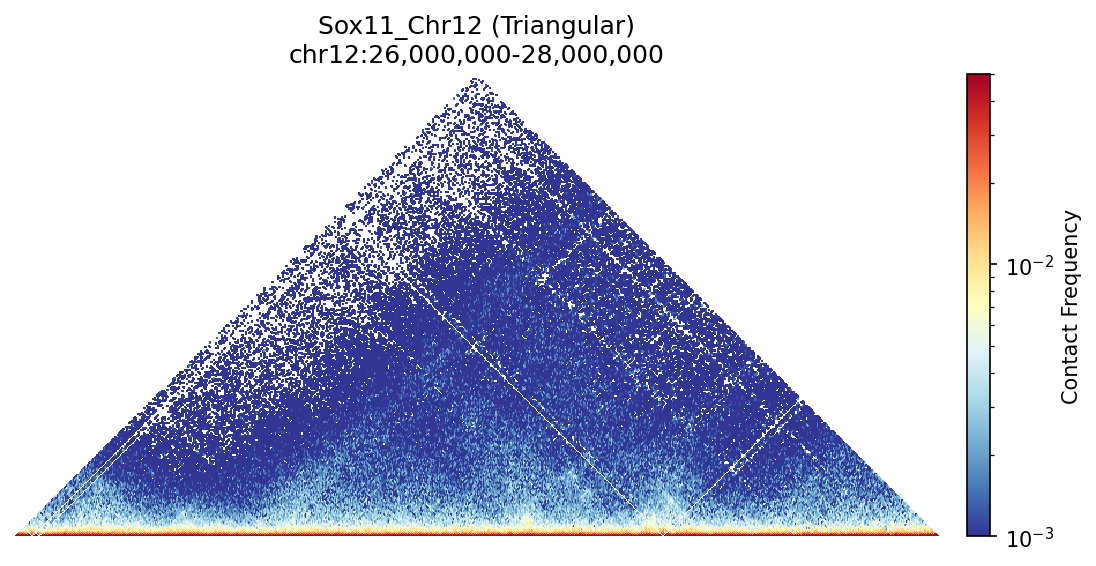
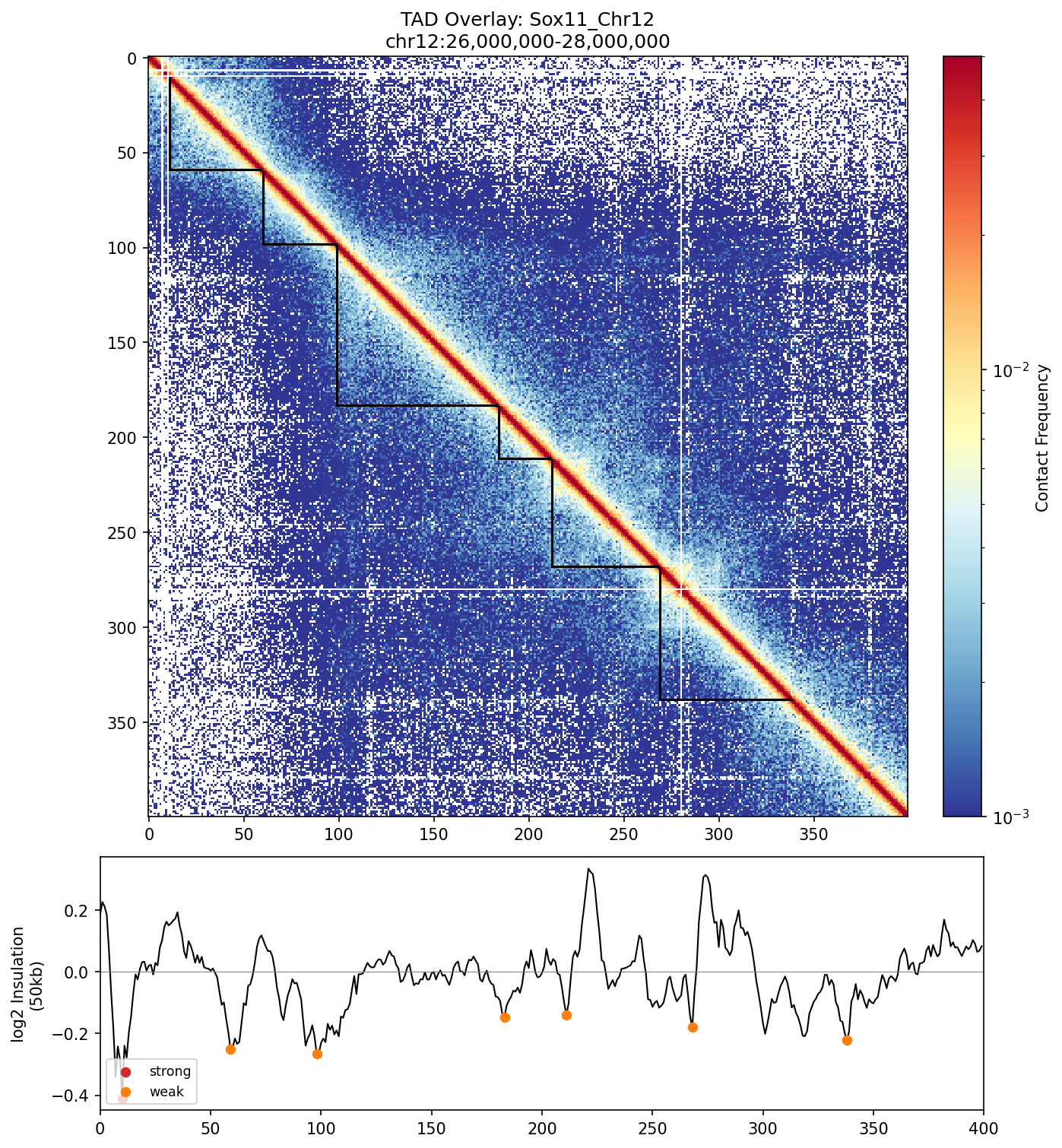
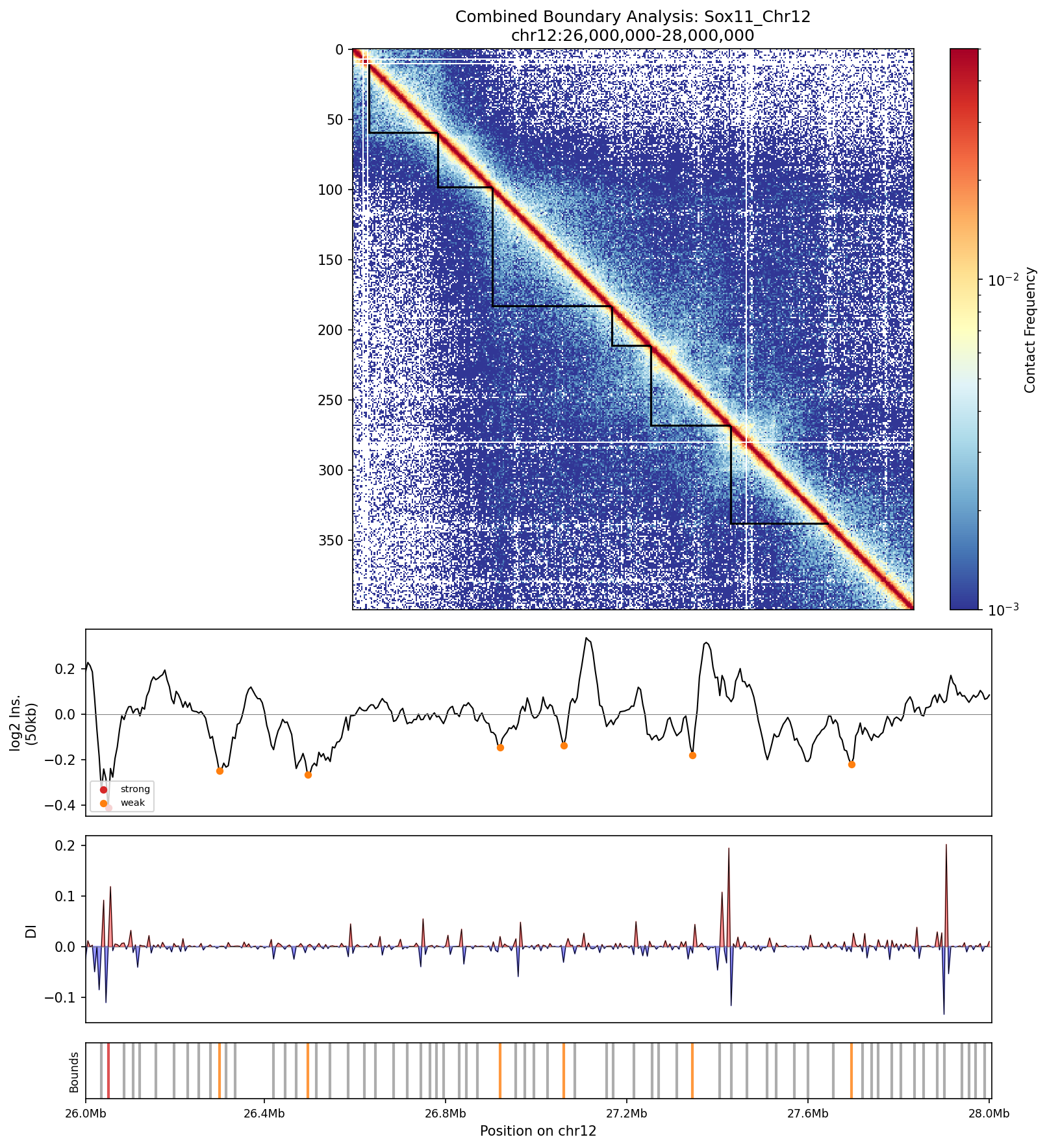
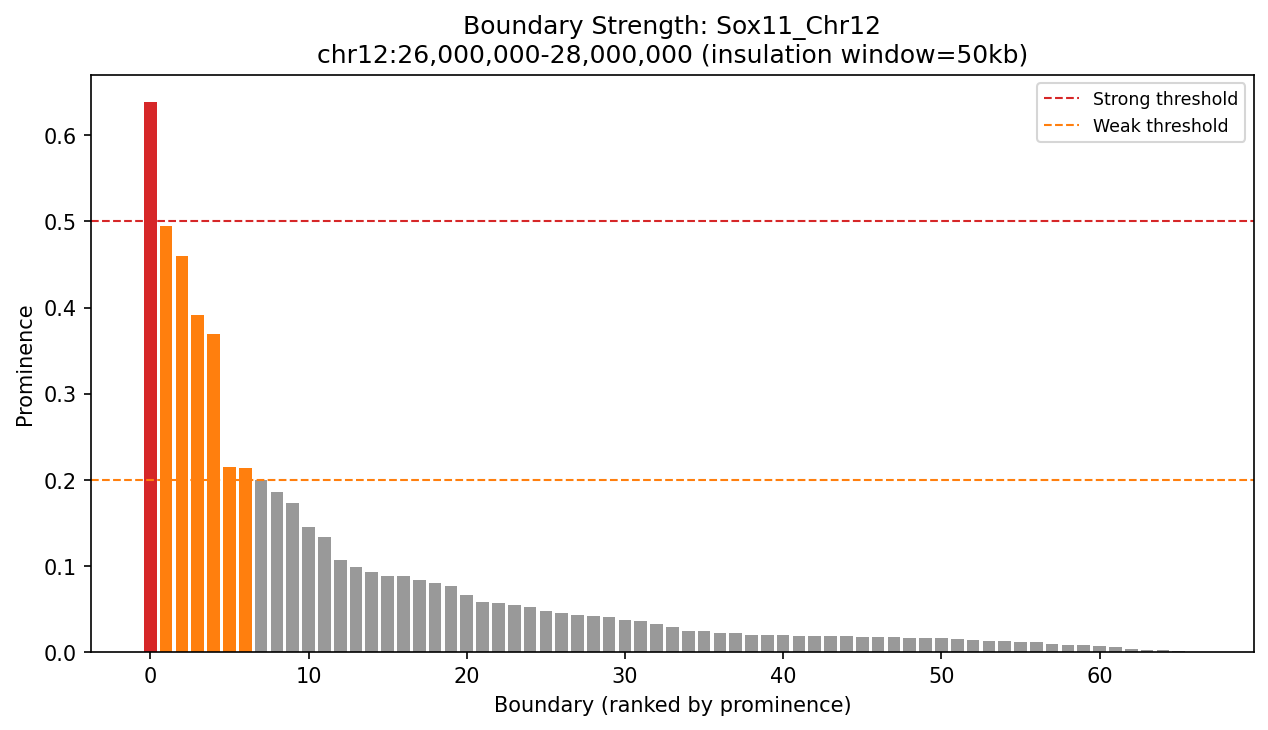
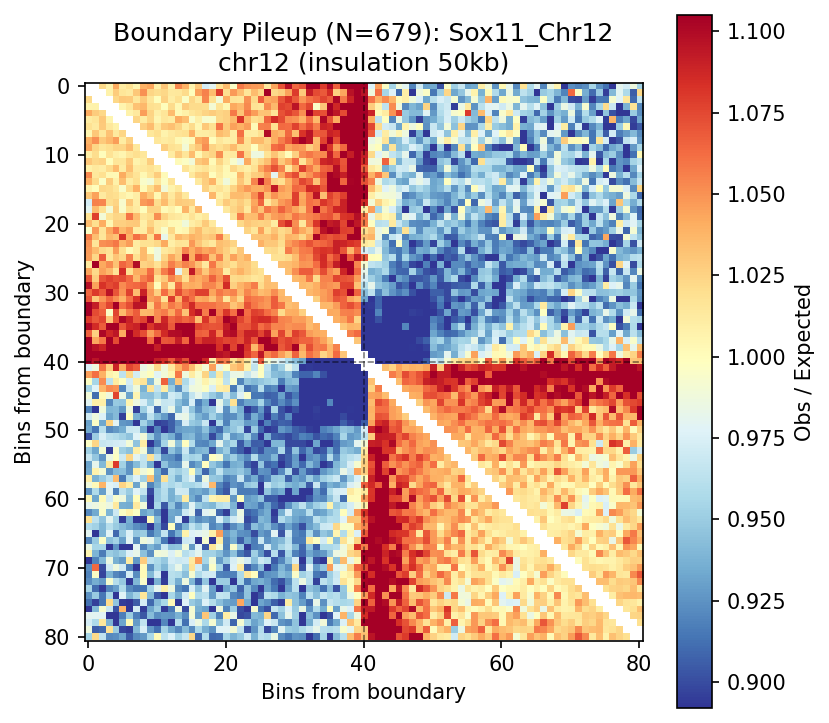
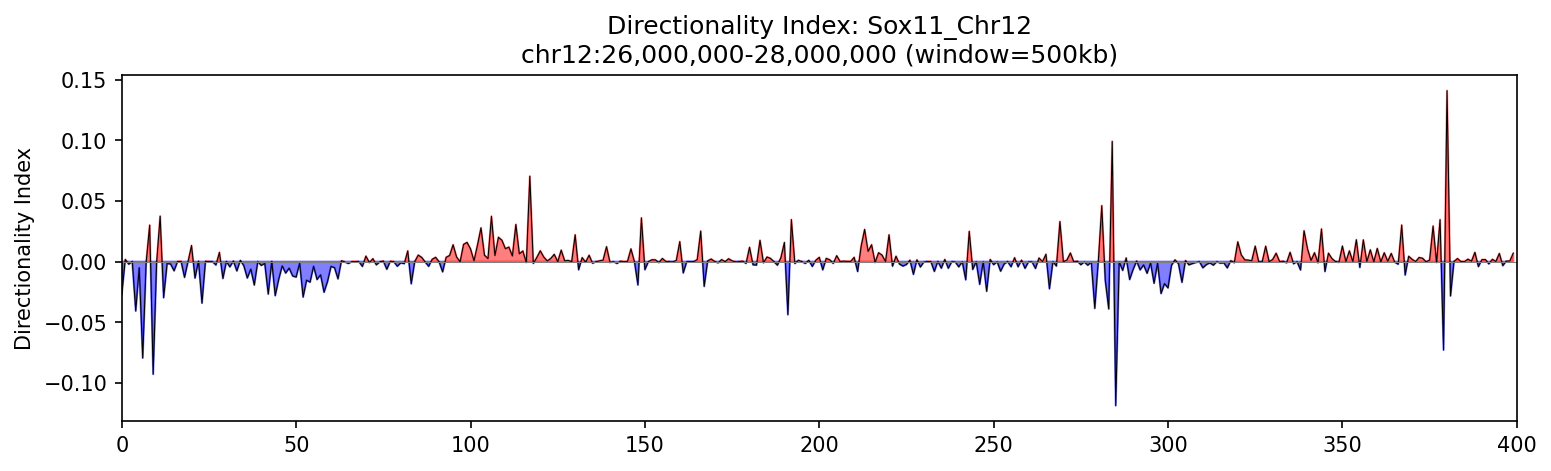
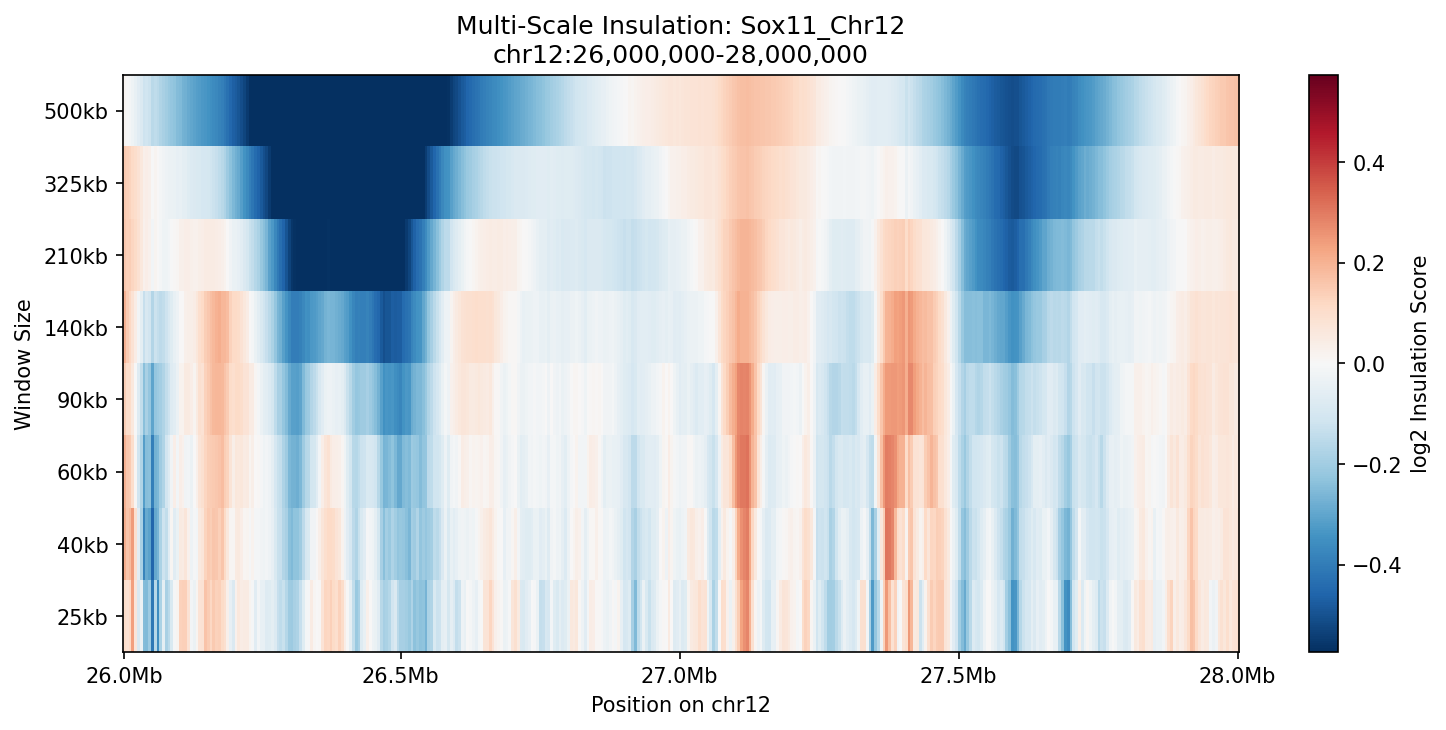
Sox11_Chr12 Region
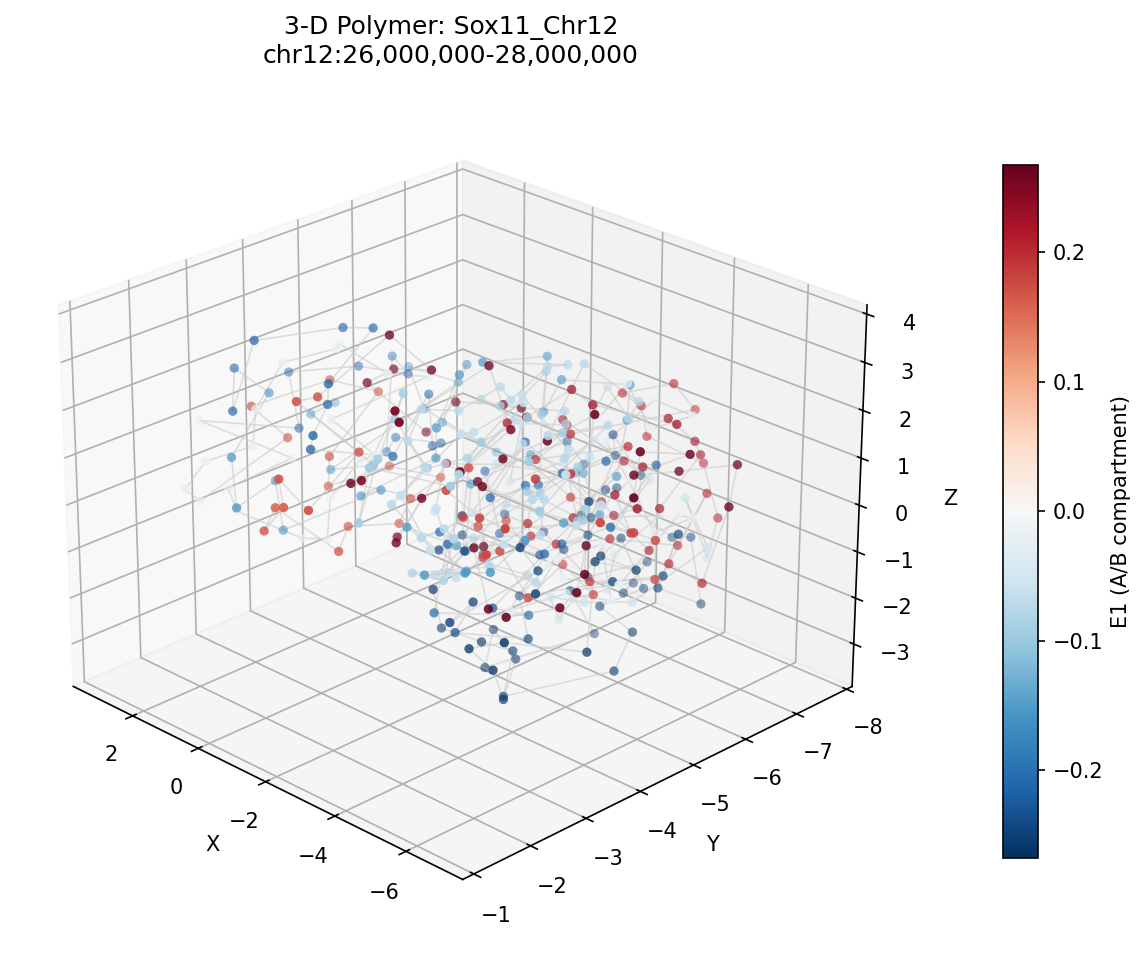
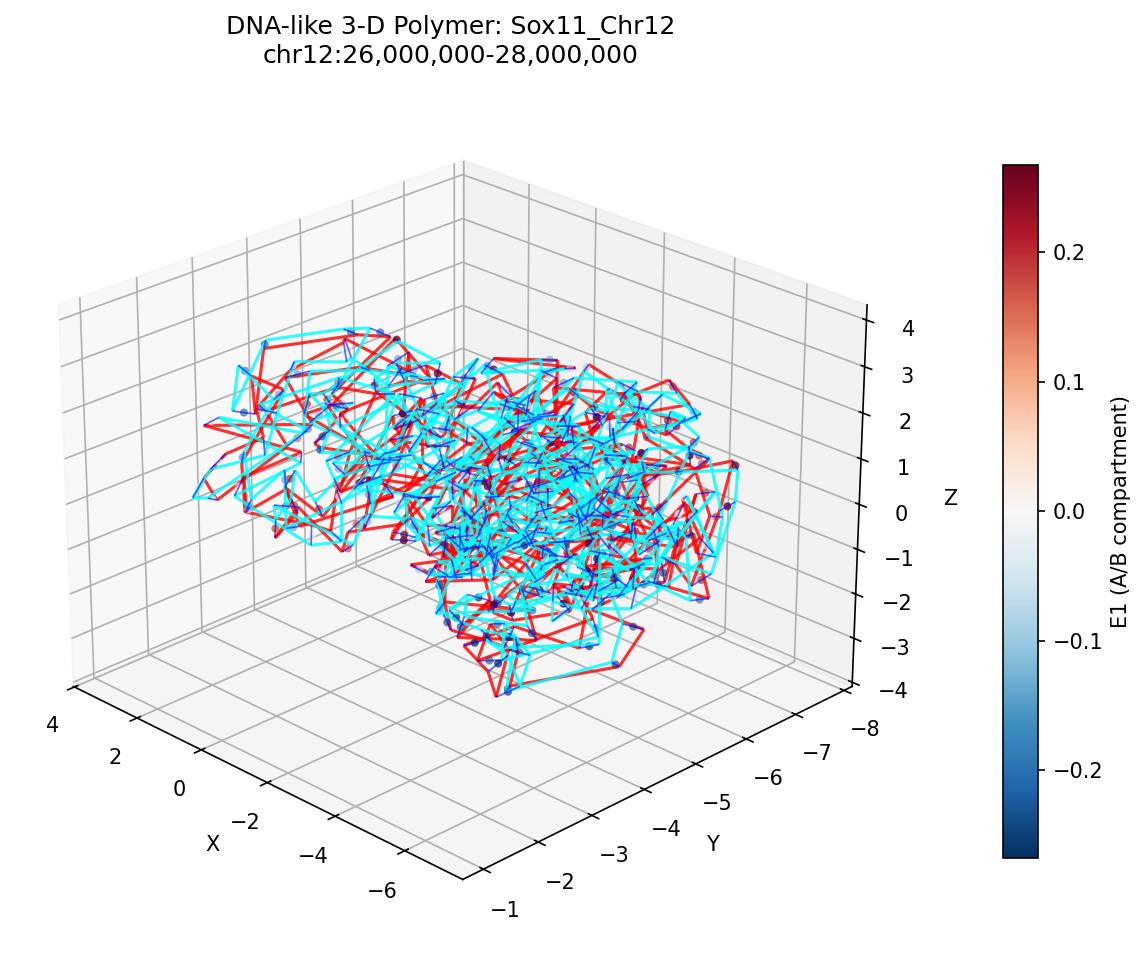
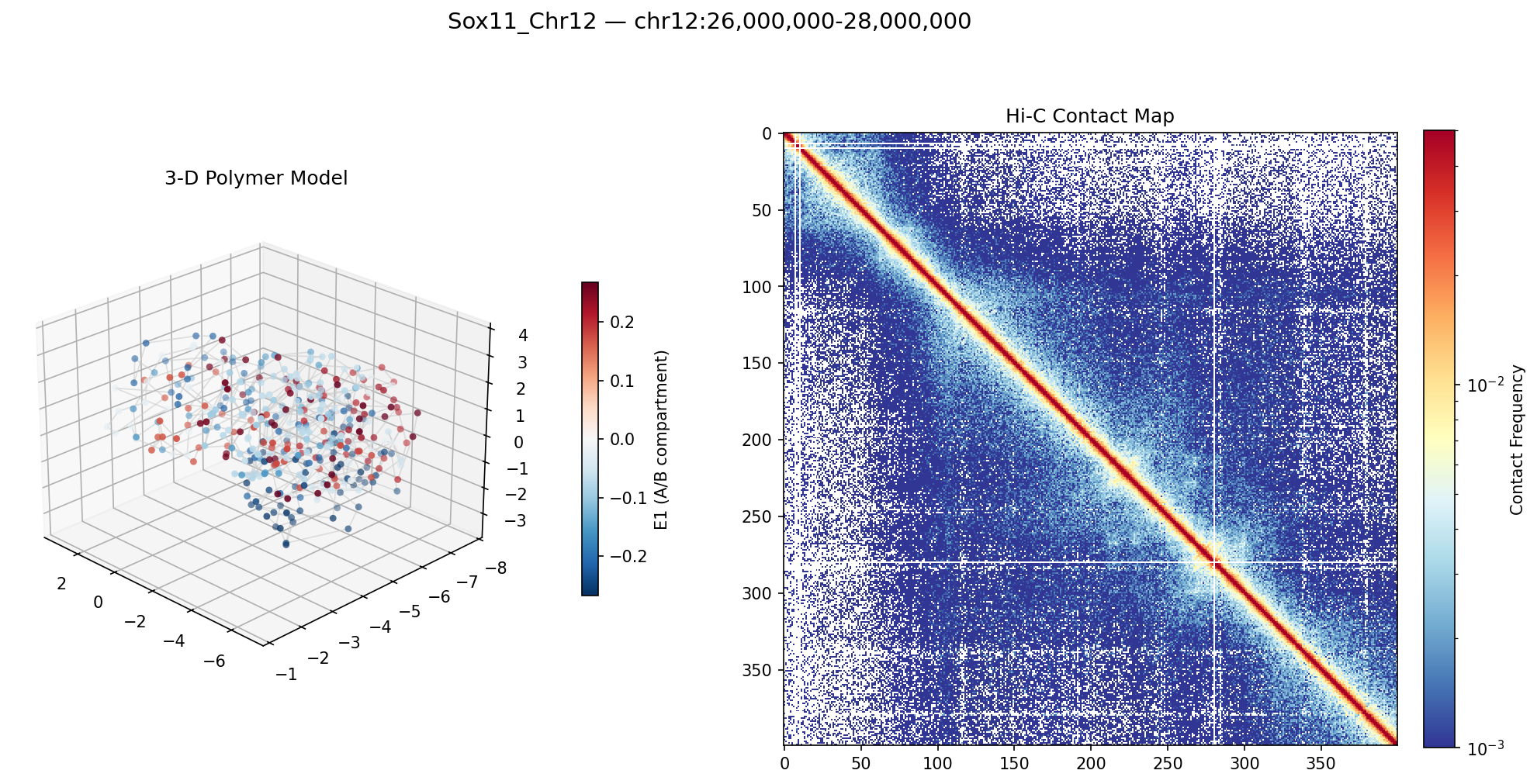
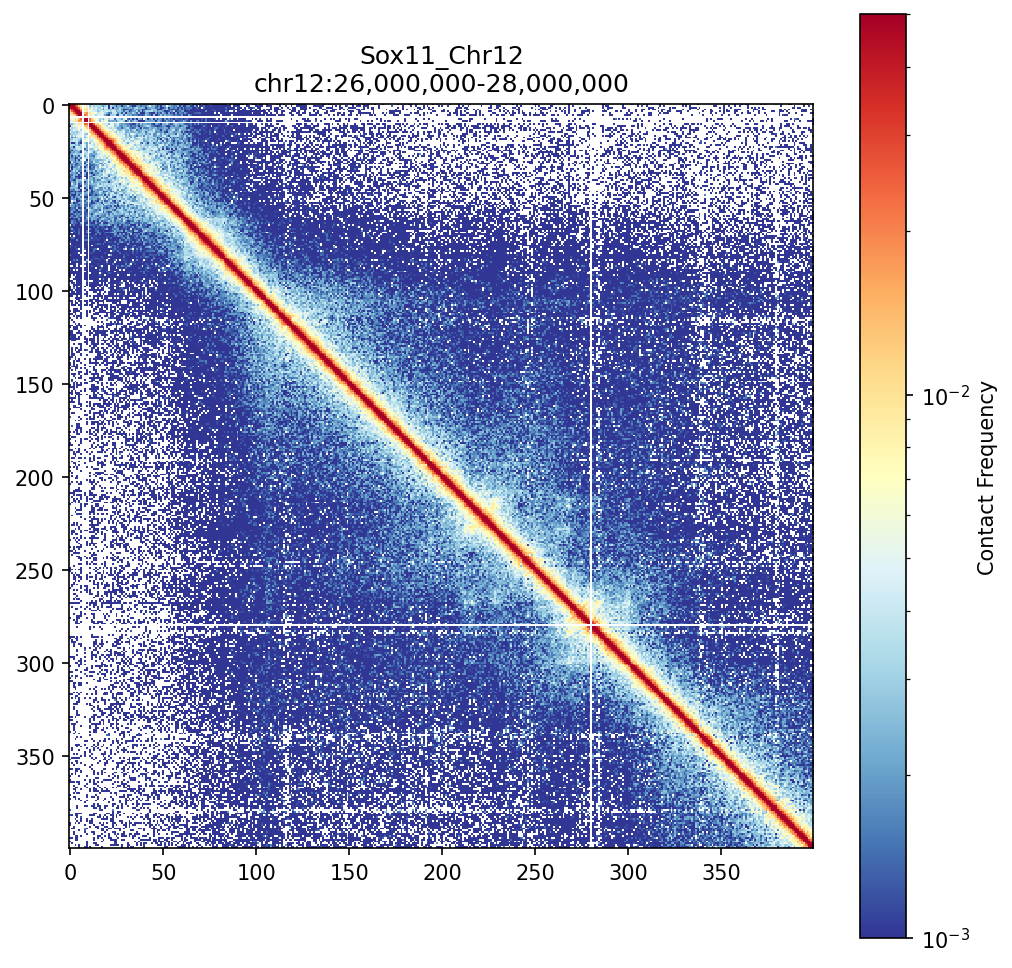
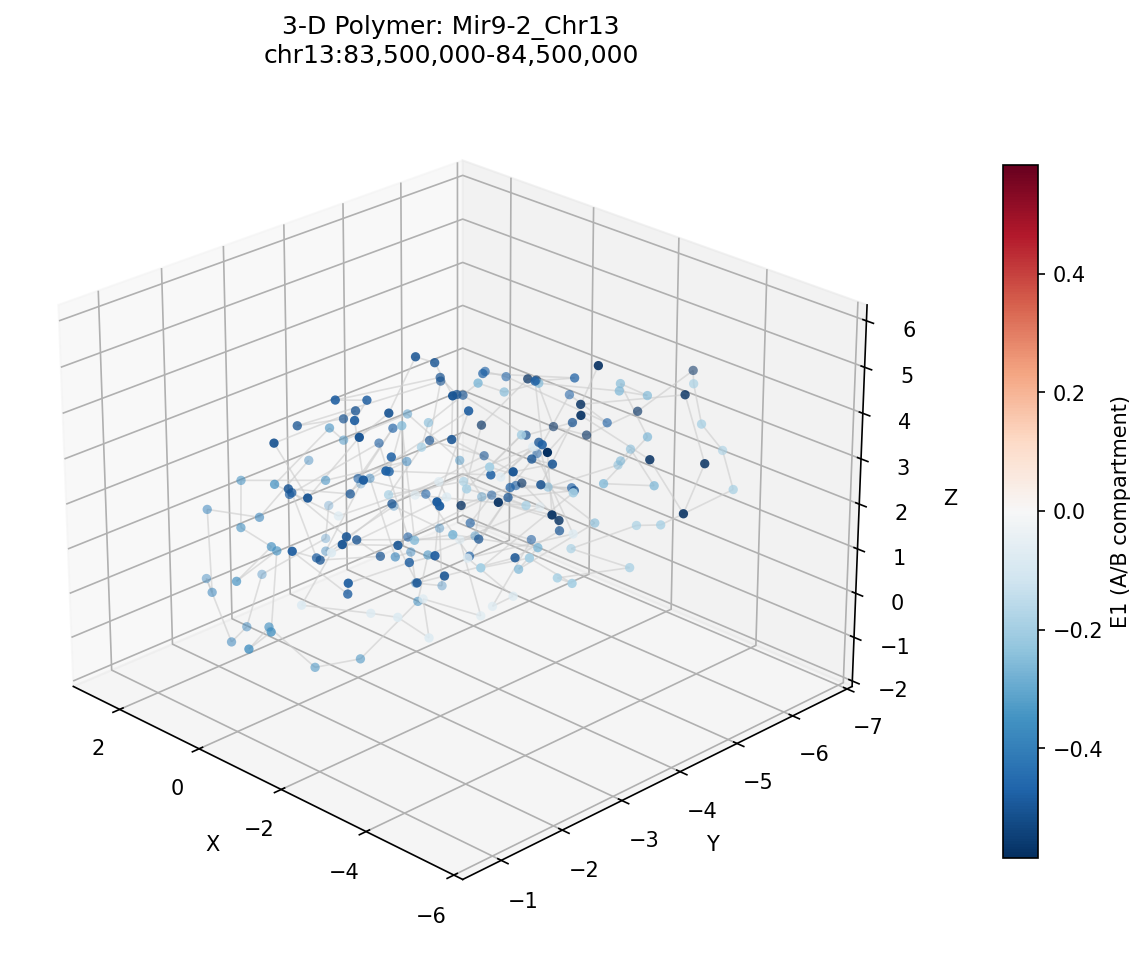
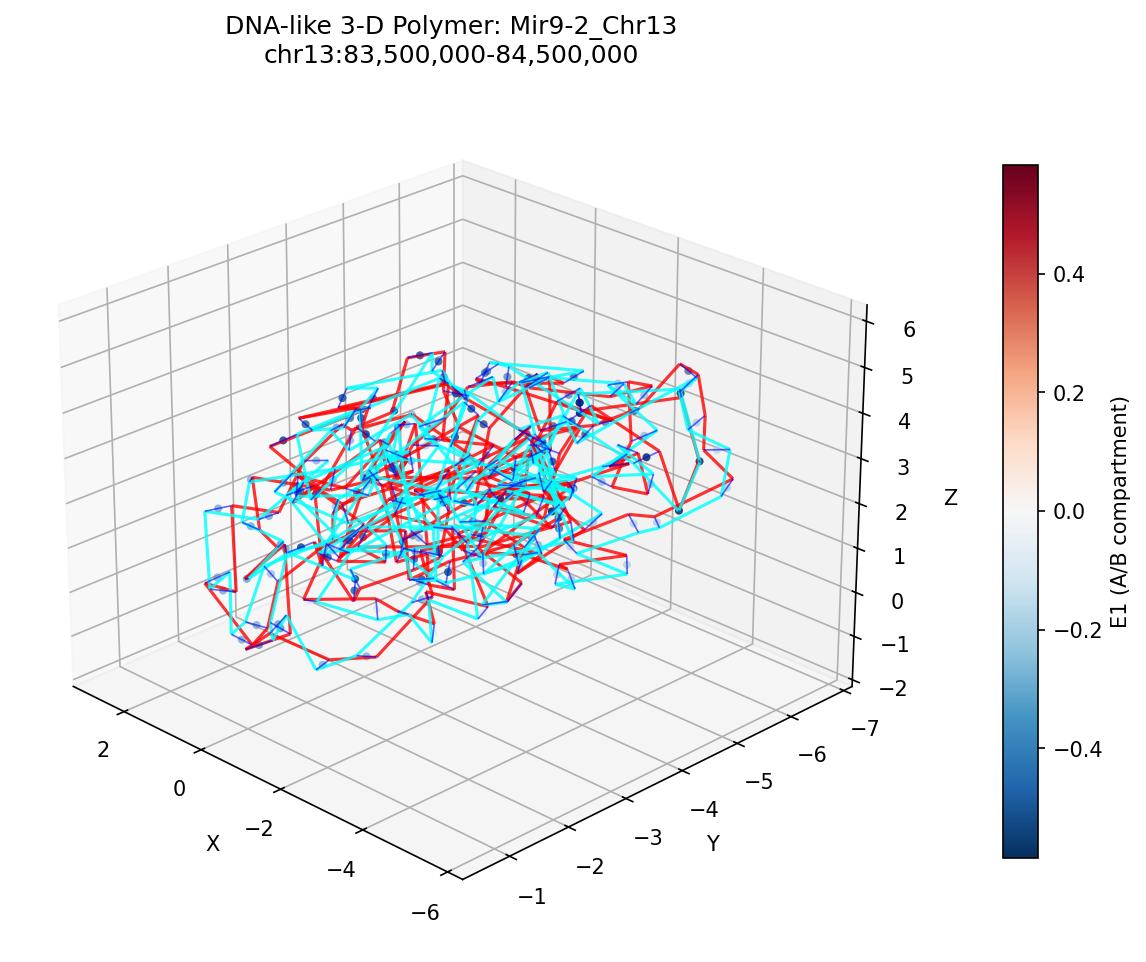
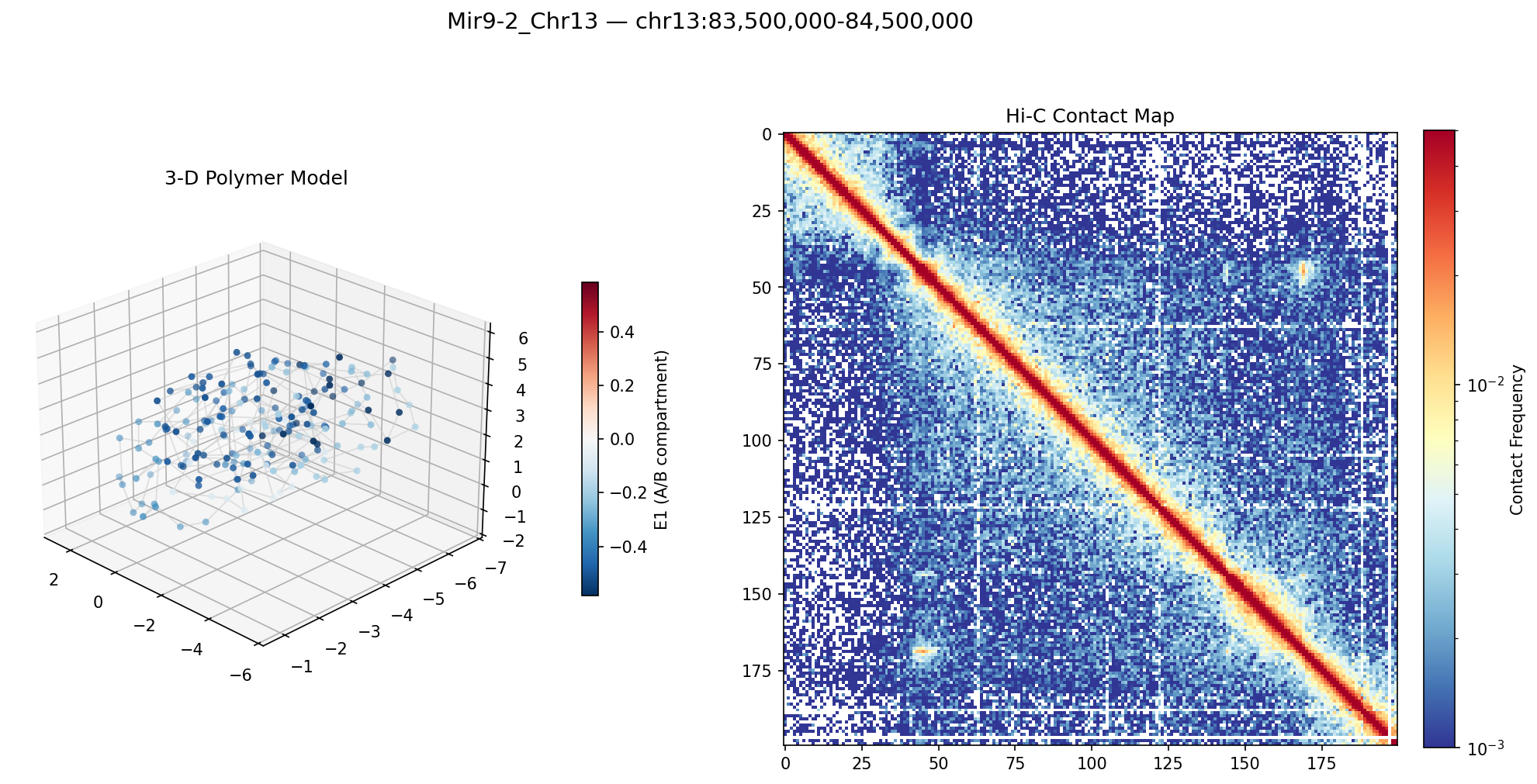
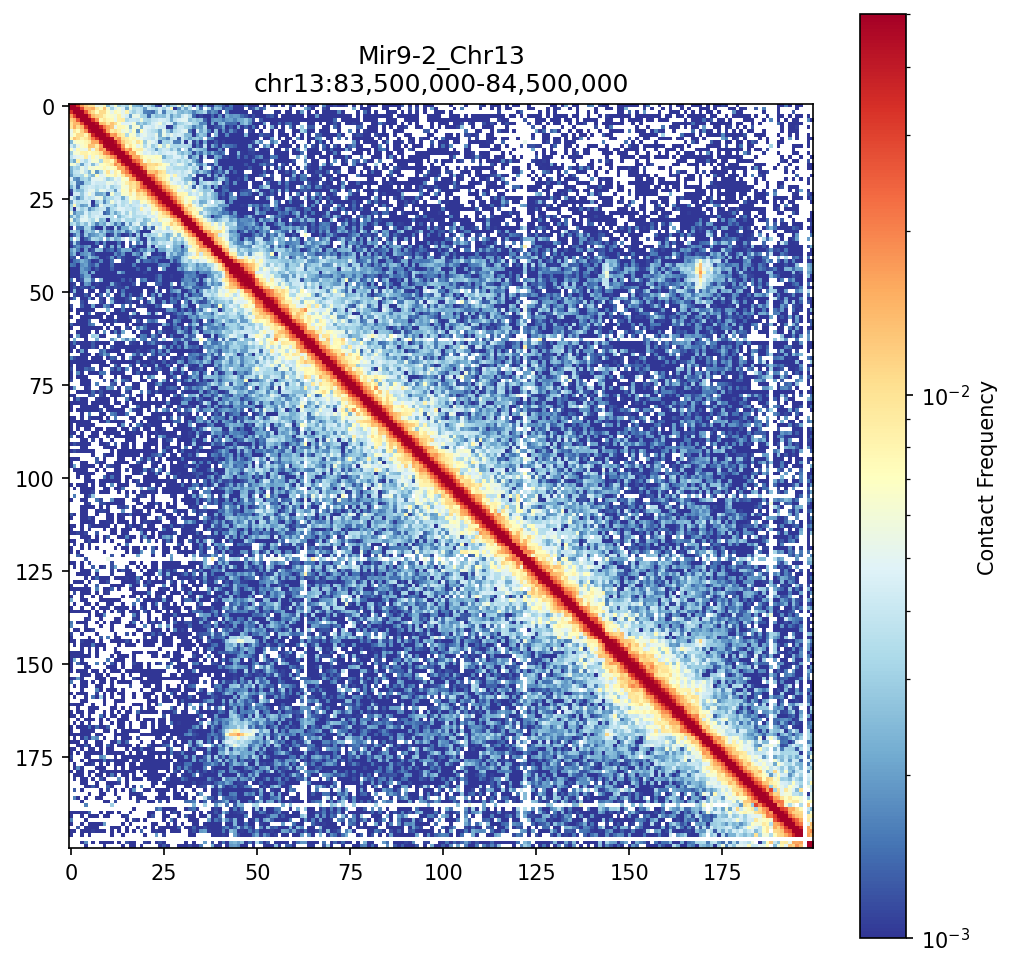
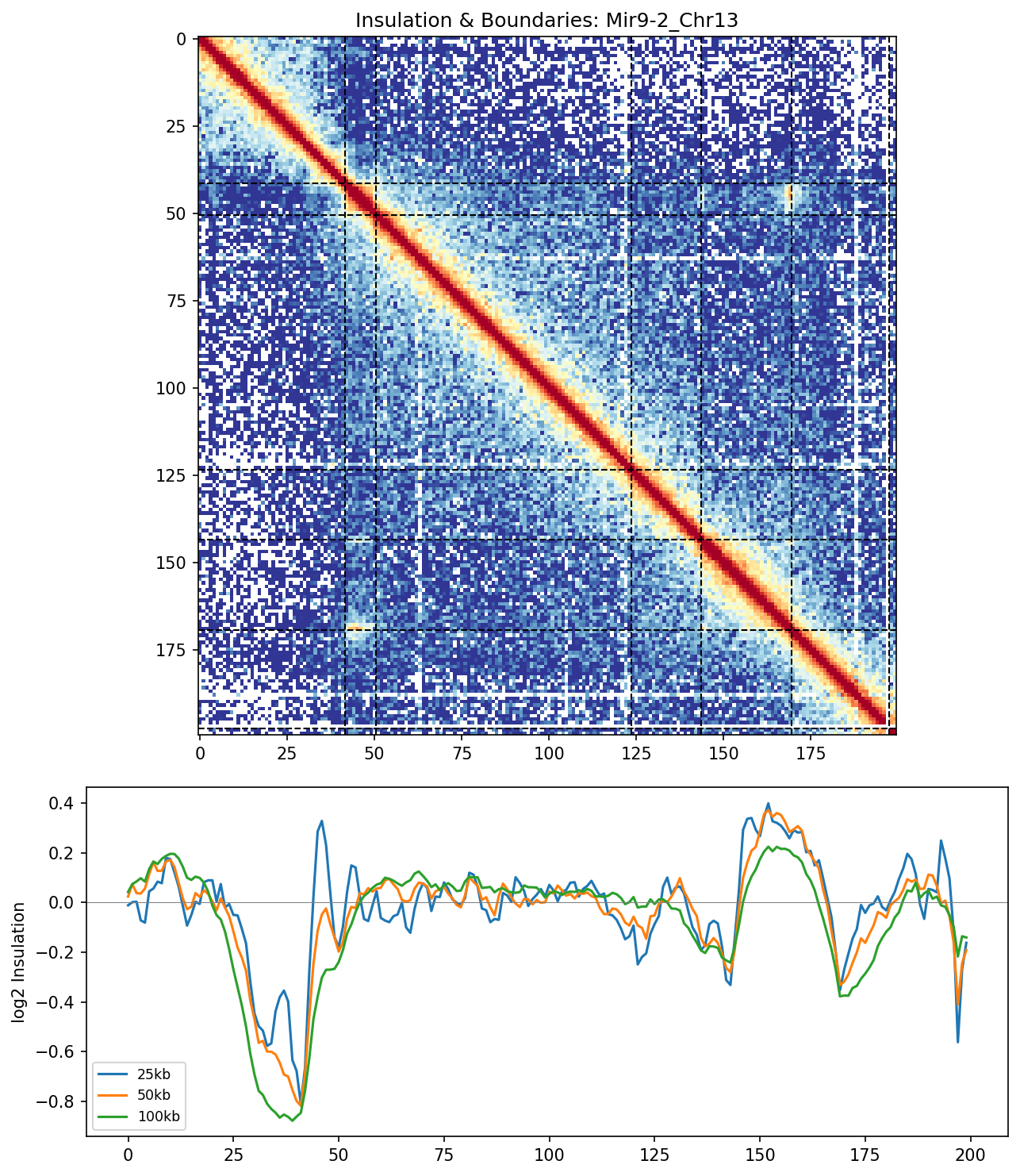
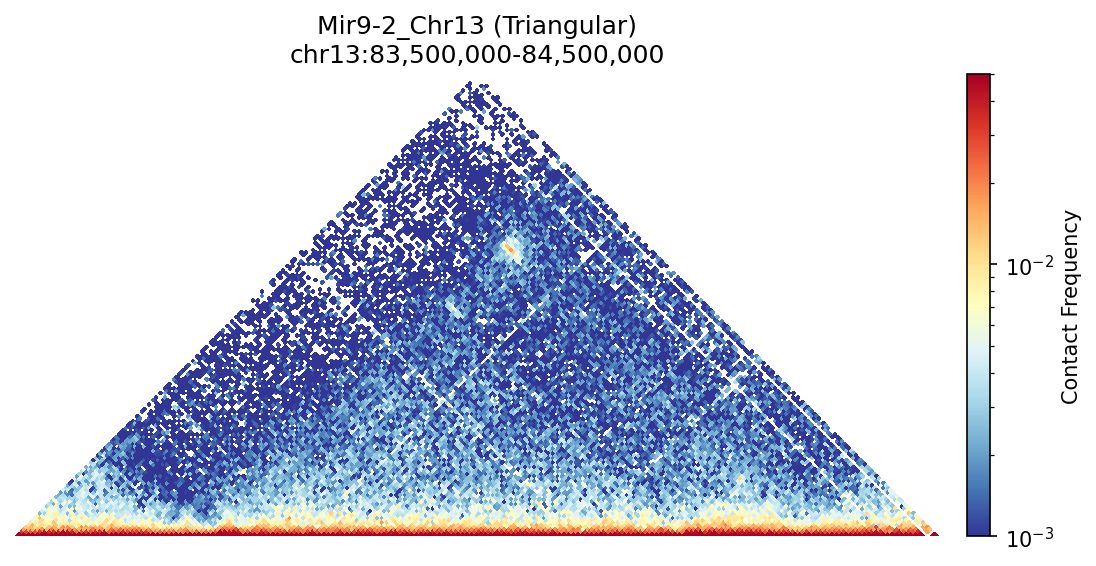
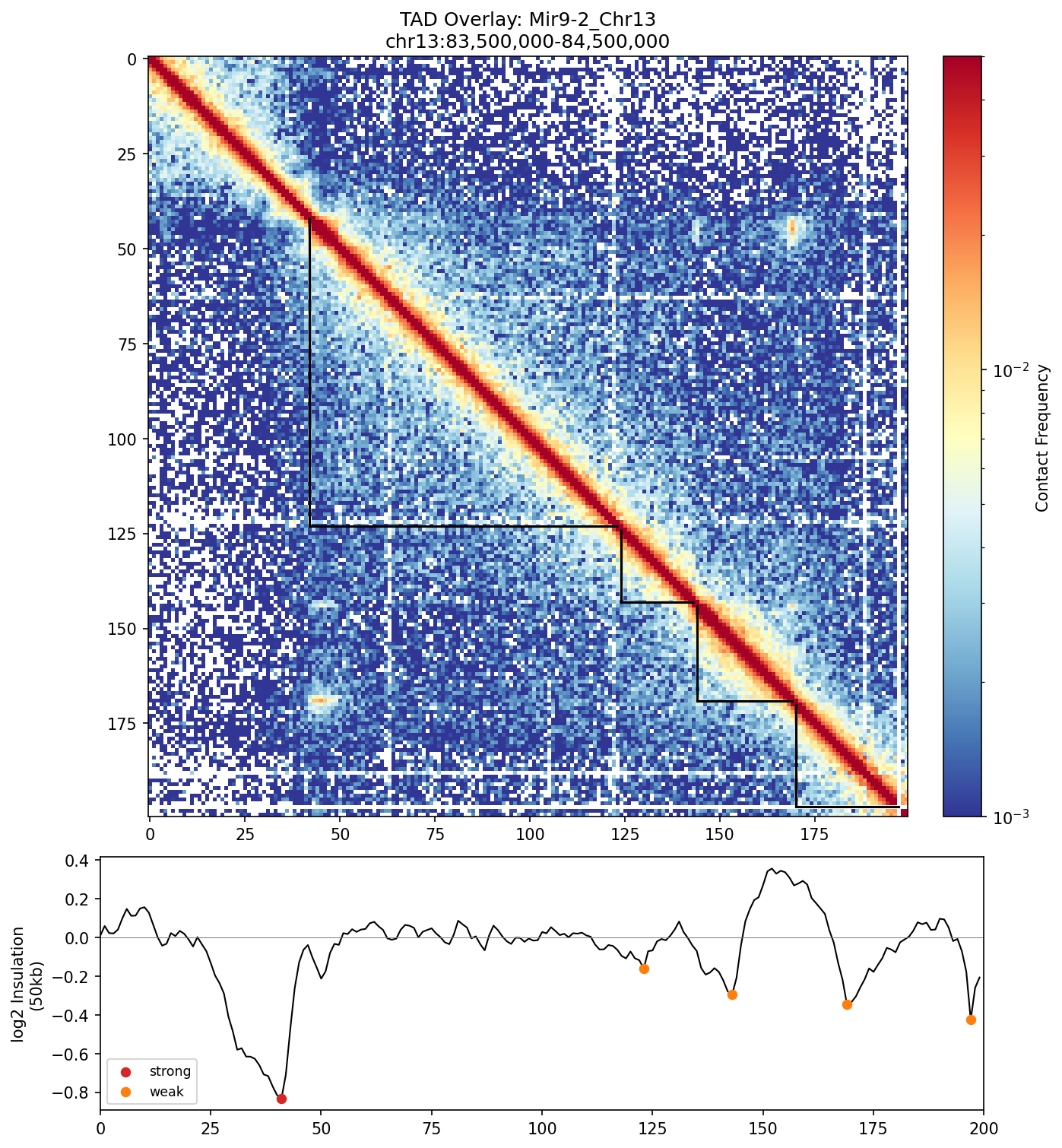
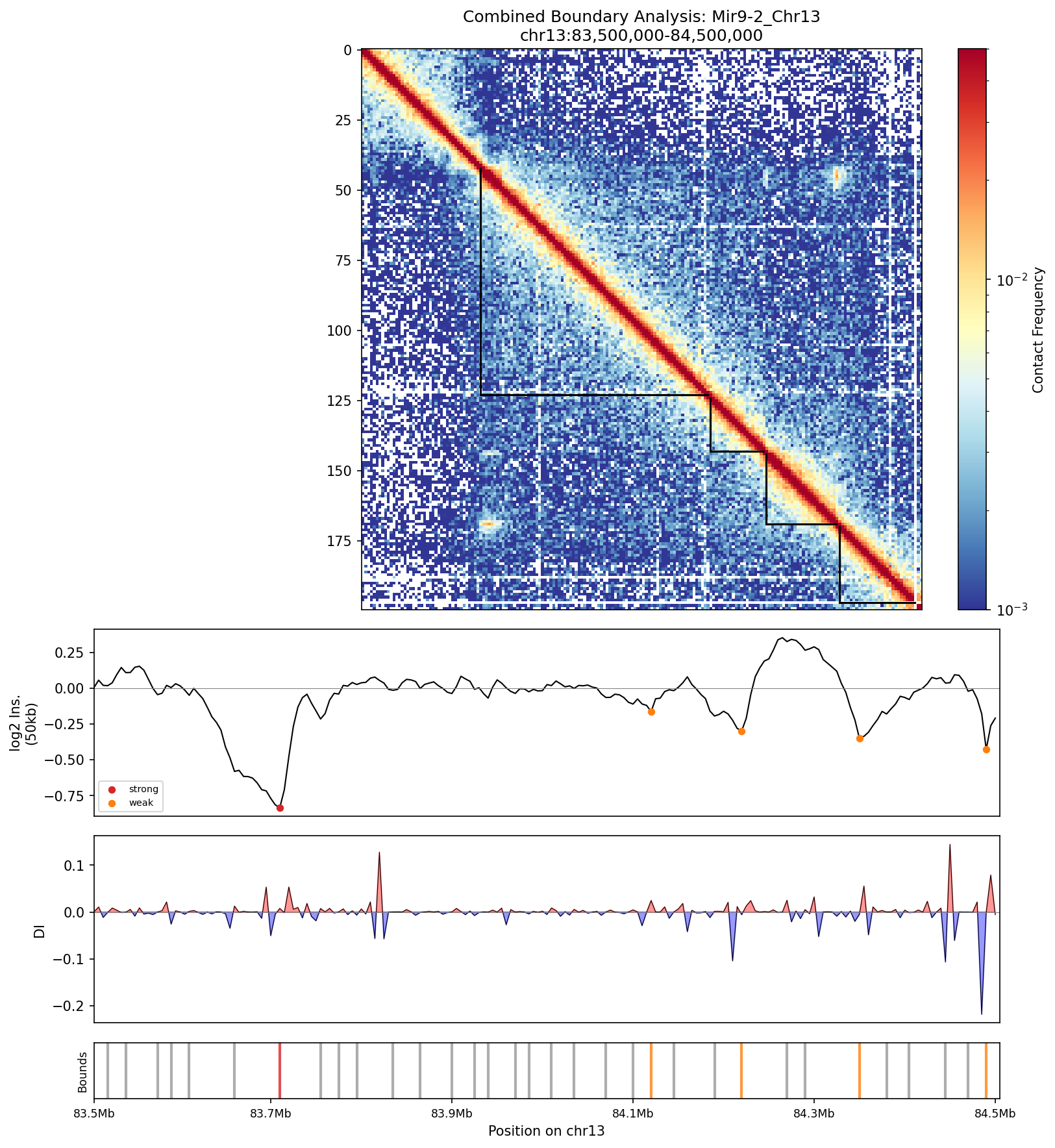
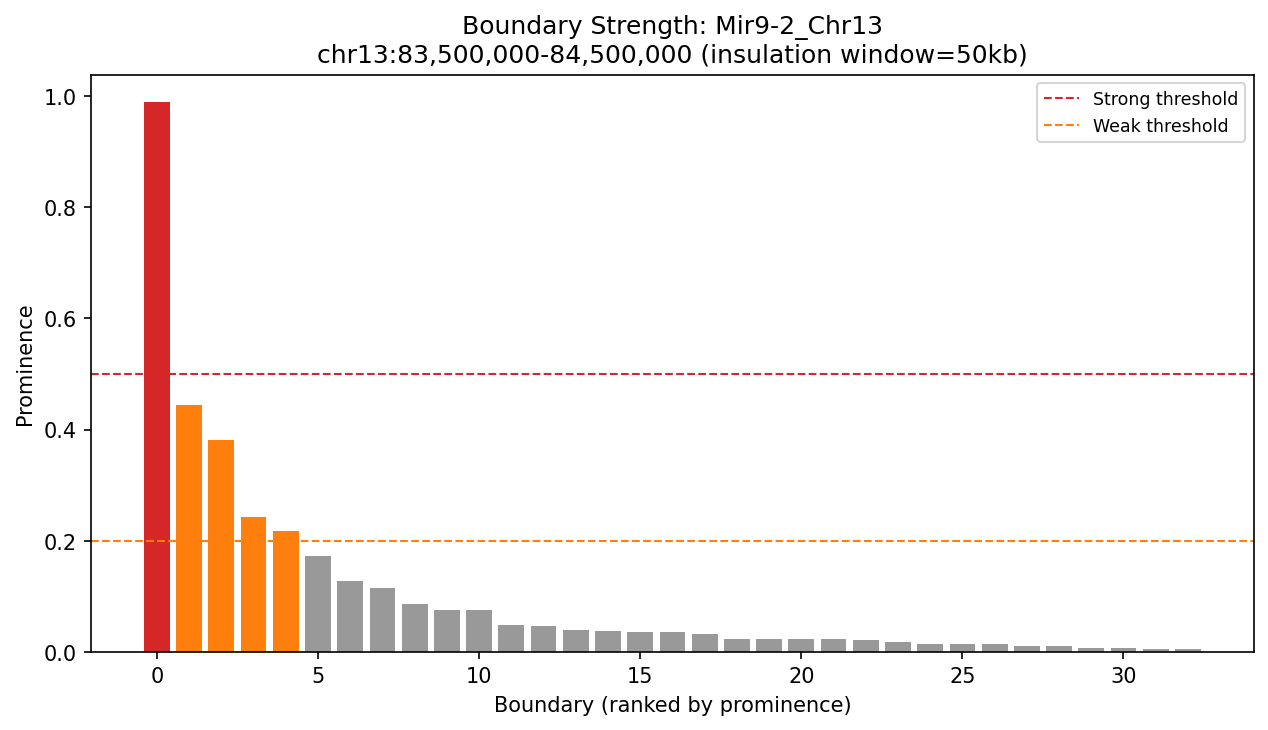
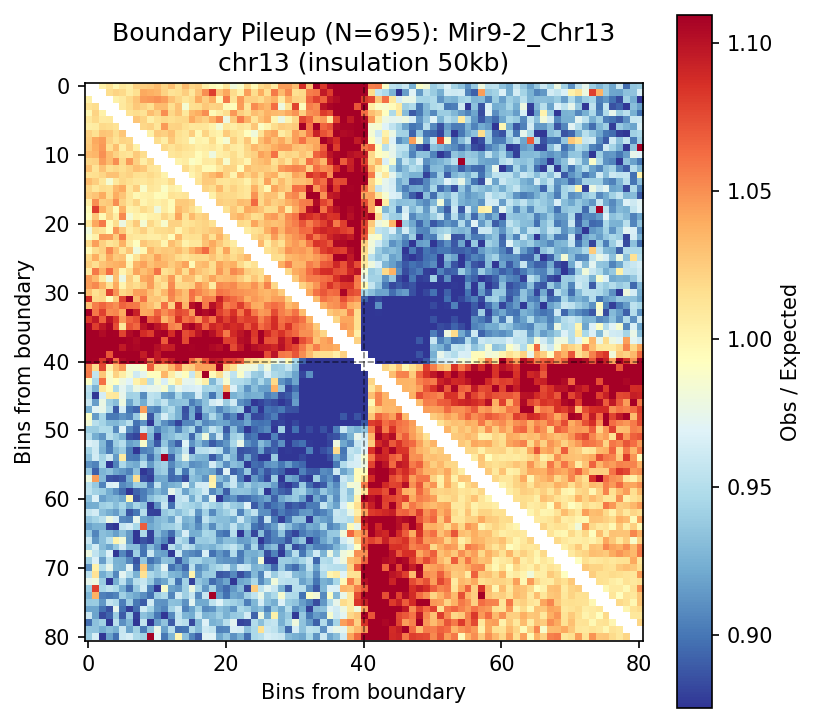
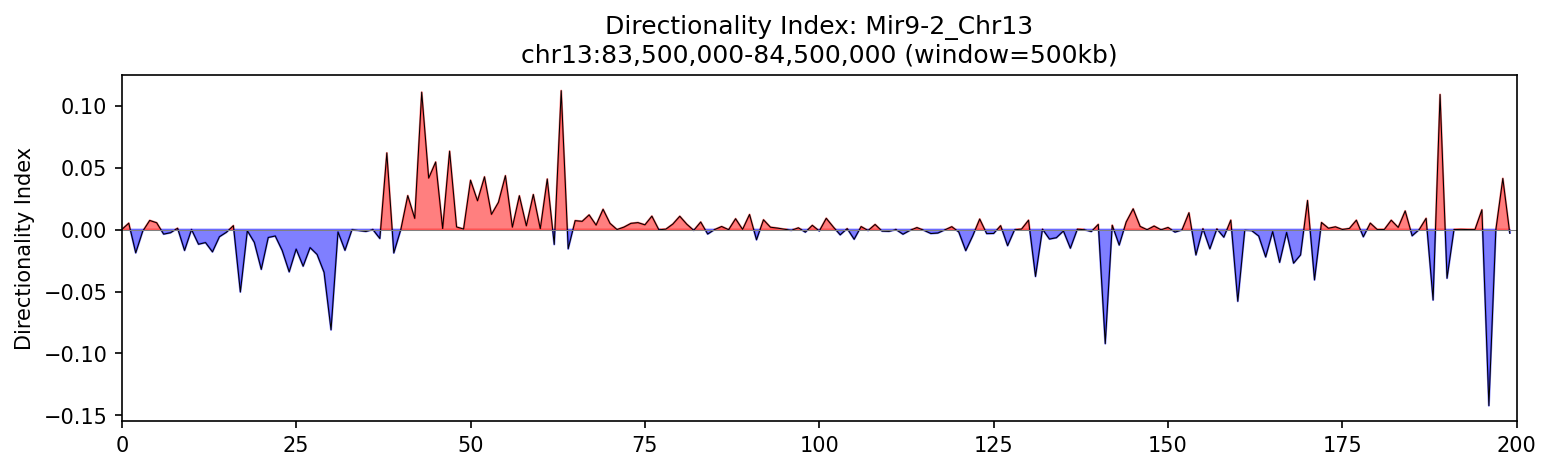
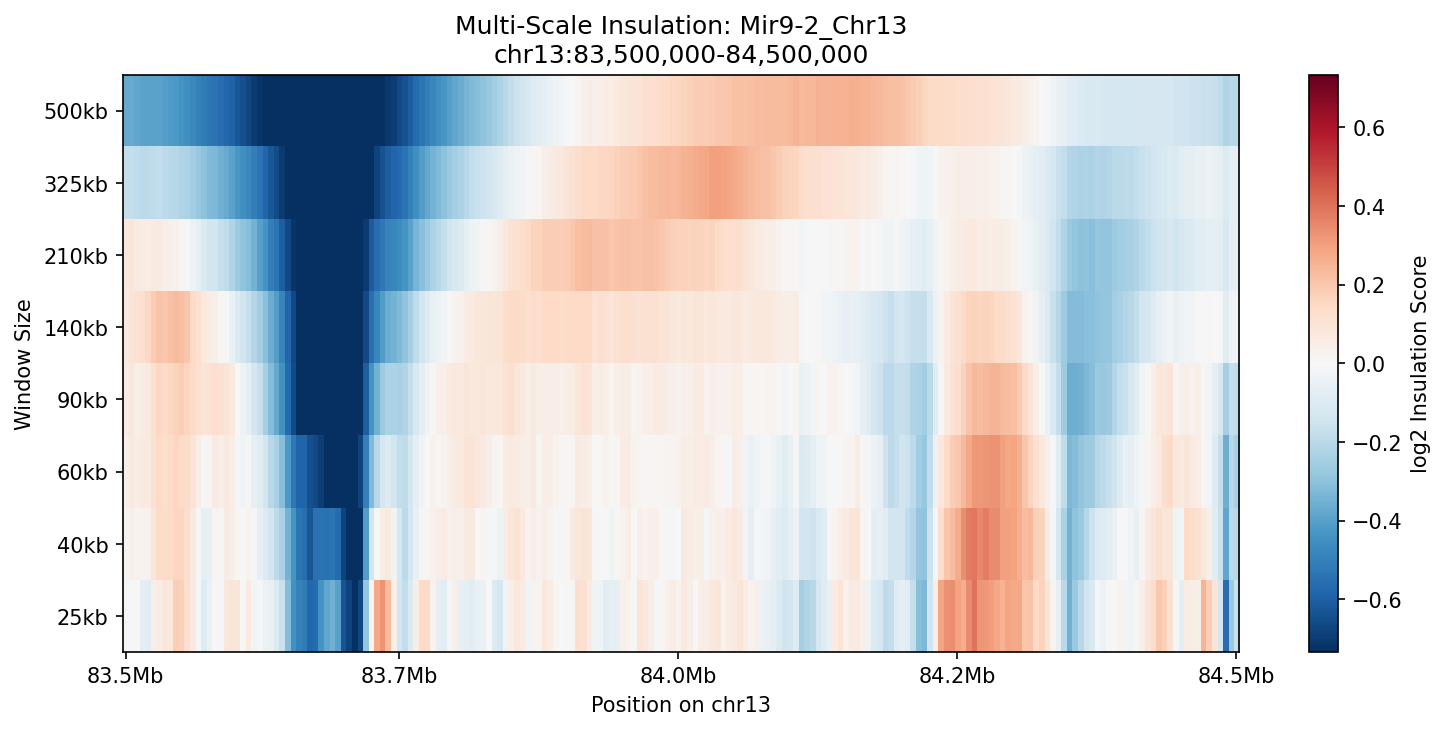
Mir9-2_Chr13 Region
Other Analyses
Dovetail® Analysis Suite NEW
Full Dovetail-aligned pipeline run on the public mouse Micro-C dataset (4DN, mm10, 5 kb). Implements all recommended Dovetail analysis branches: loops, compartments, HiChIP, Capture Hi-C, SVs, CNVs, and phasing. → Full Report
Chromatin Loops (Donut Background Model)
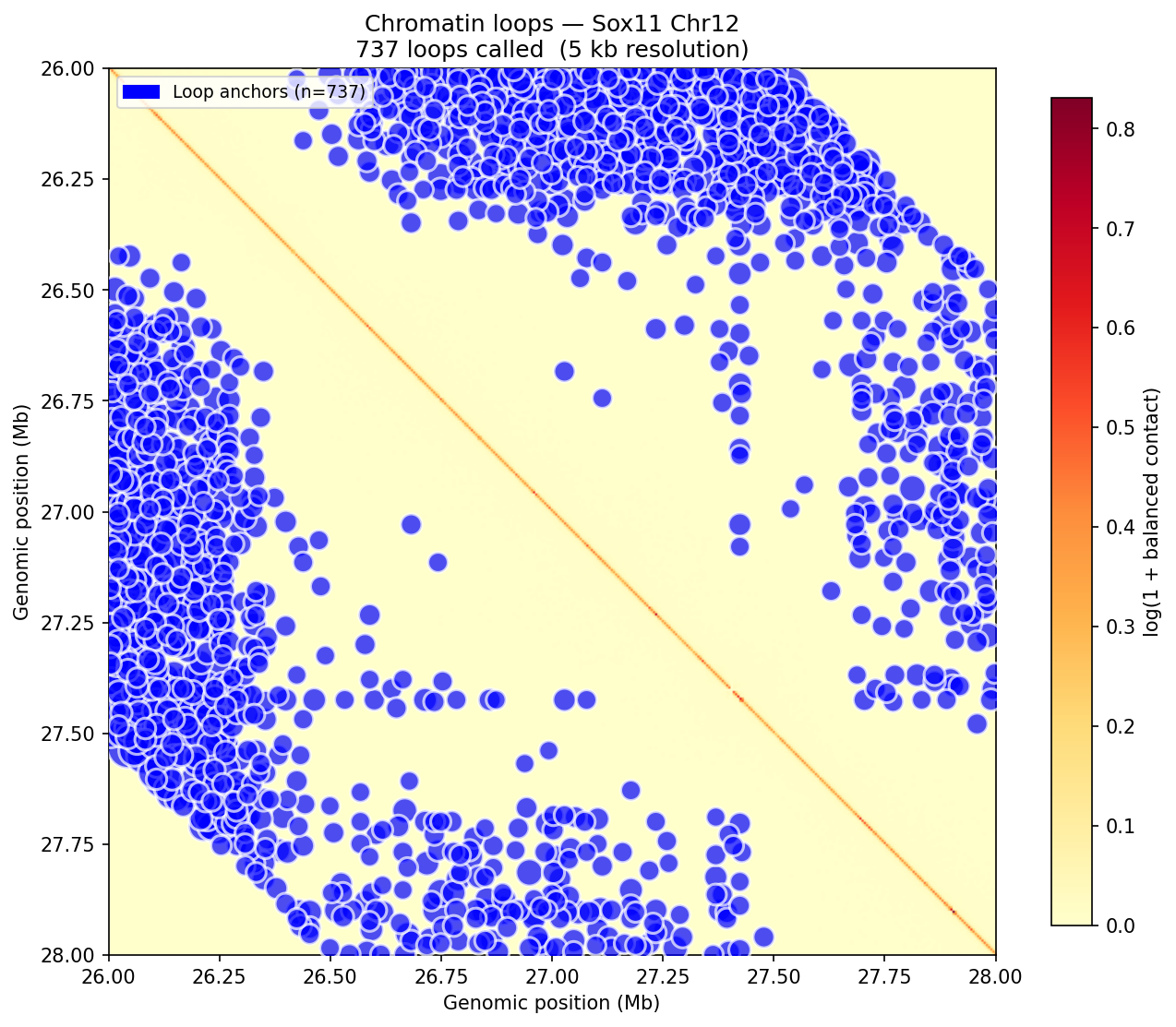
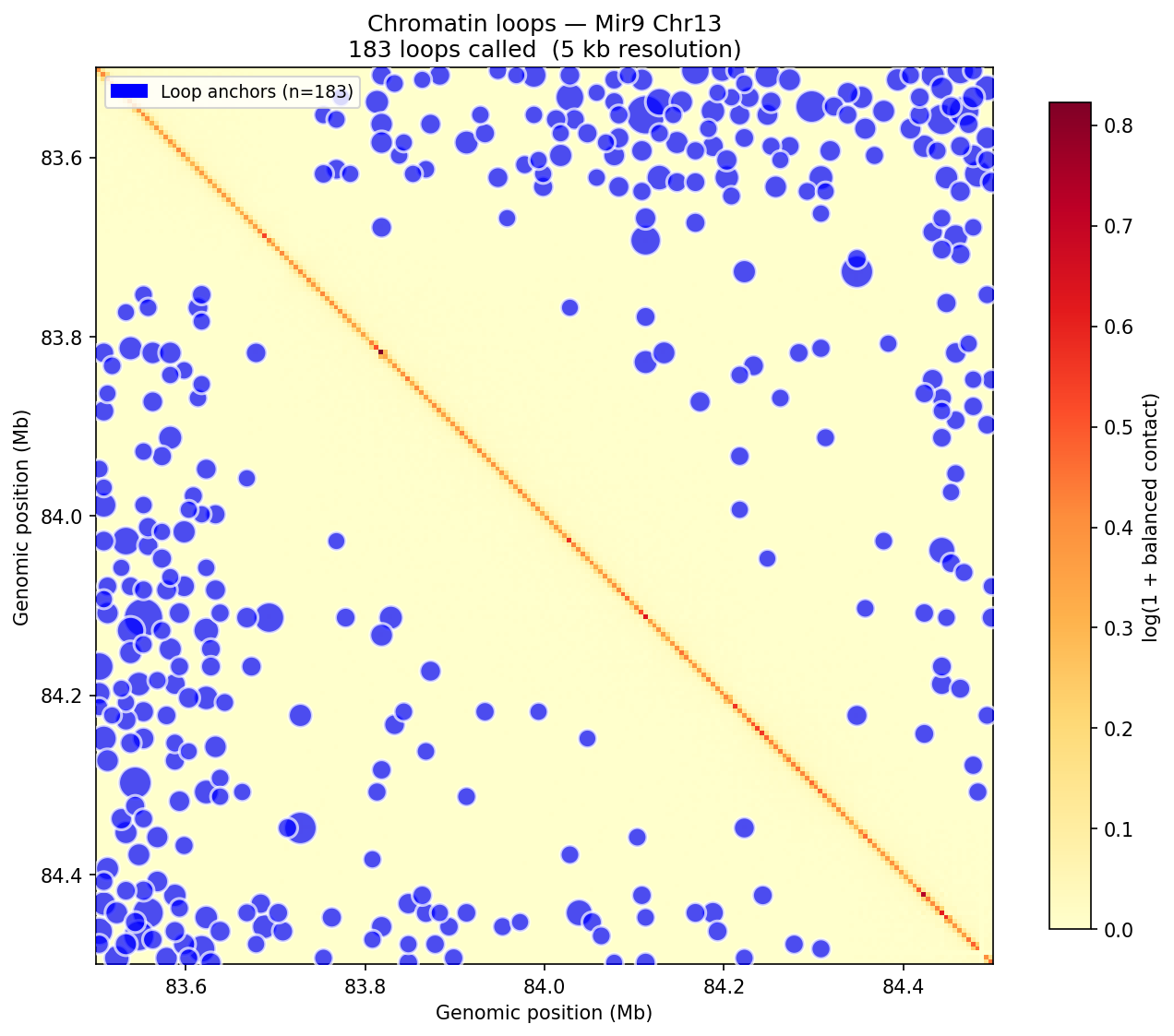
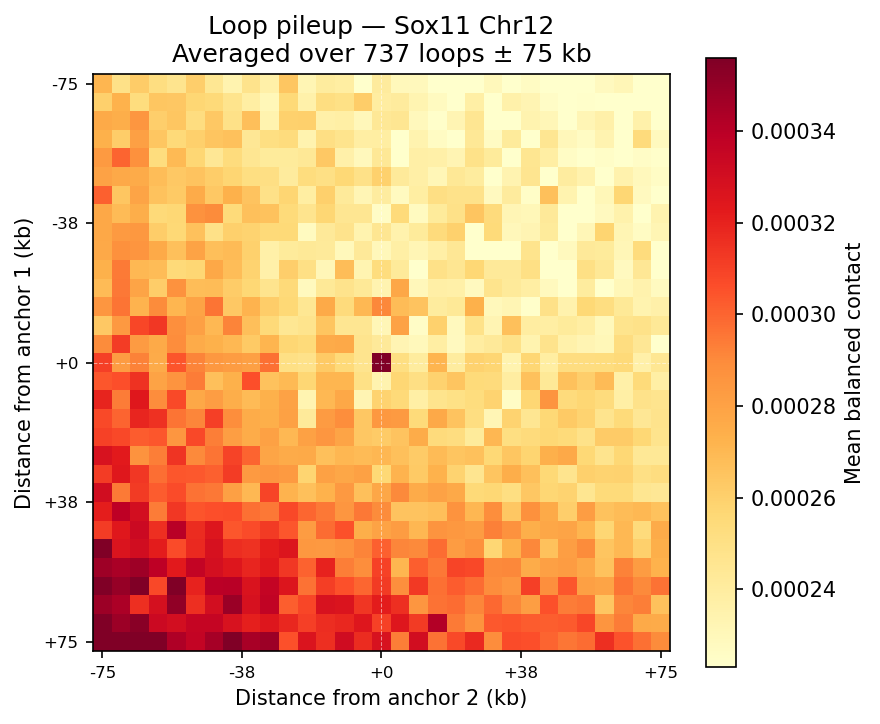
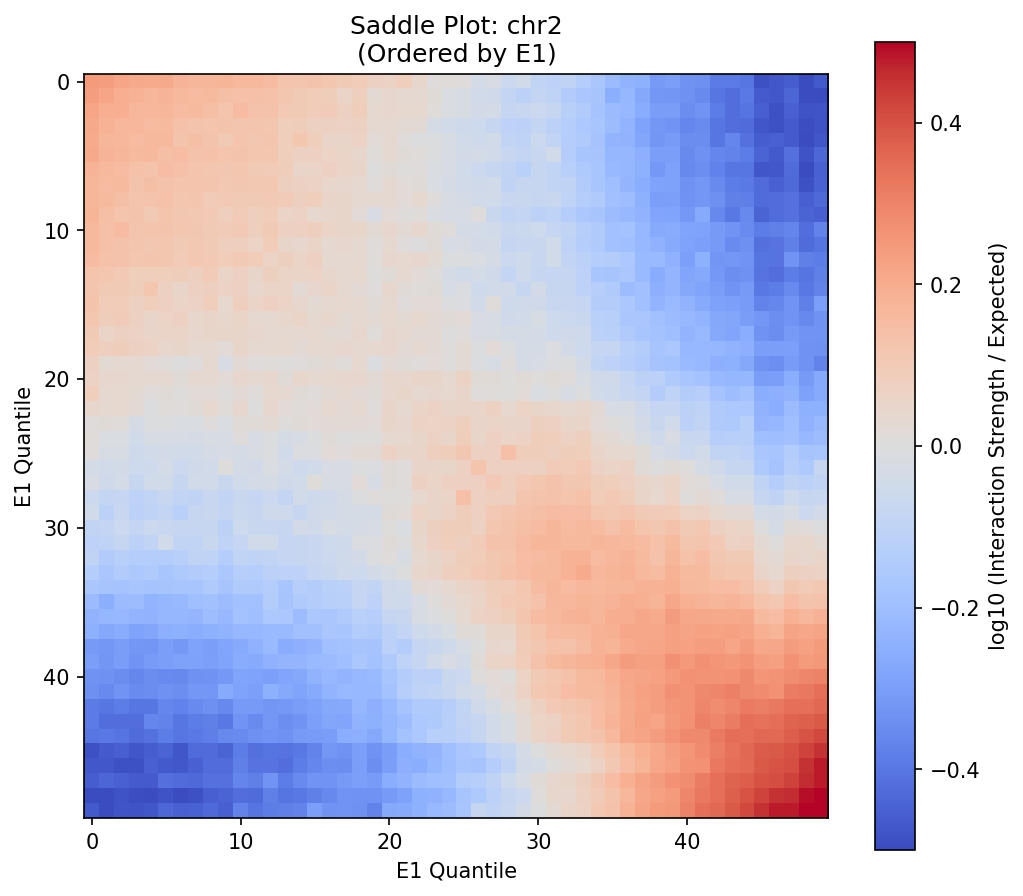
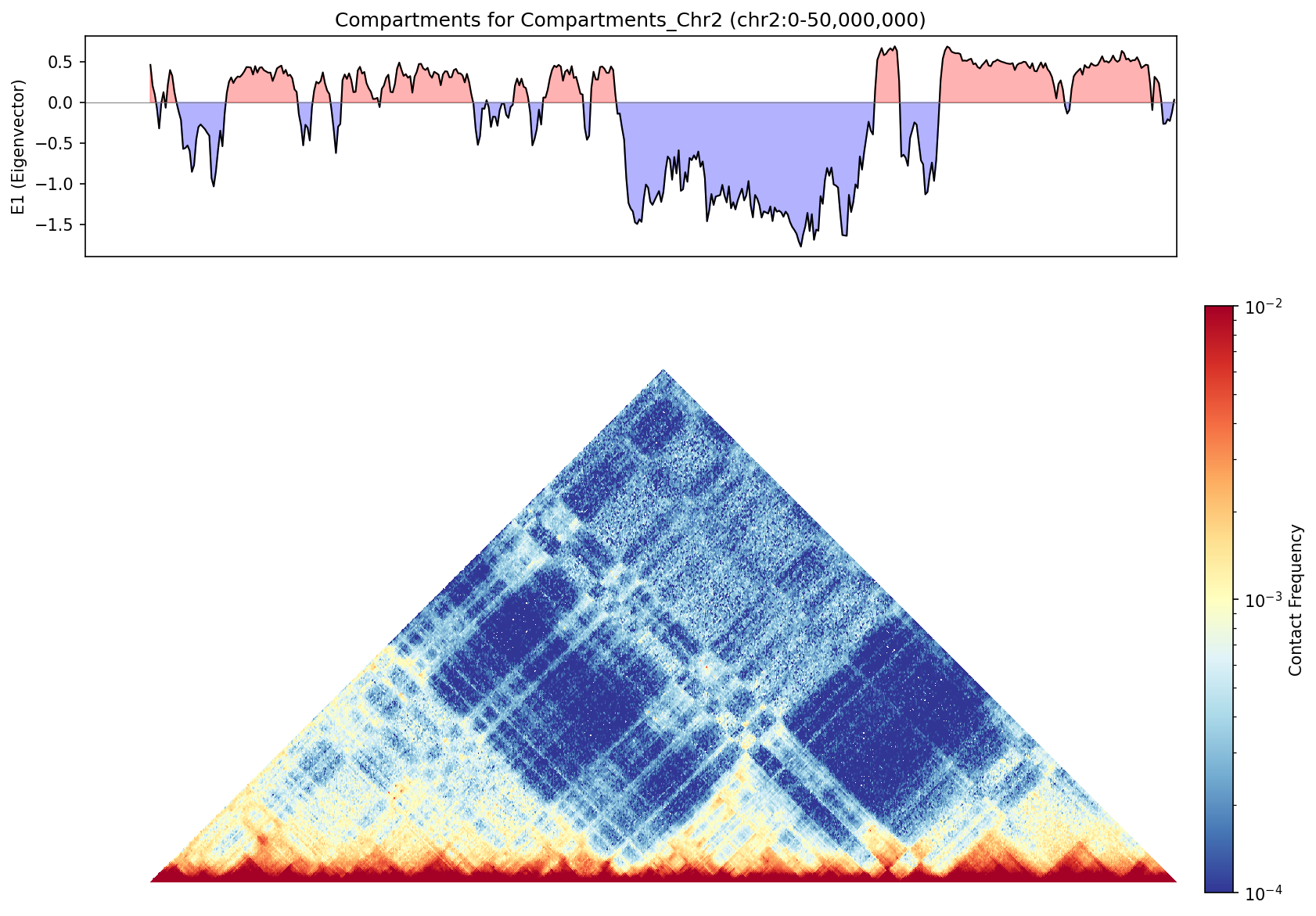
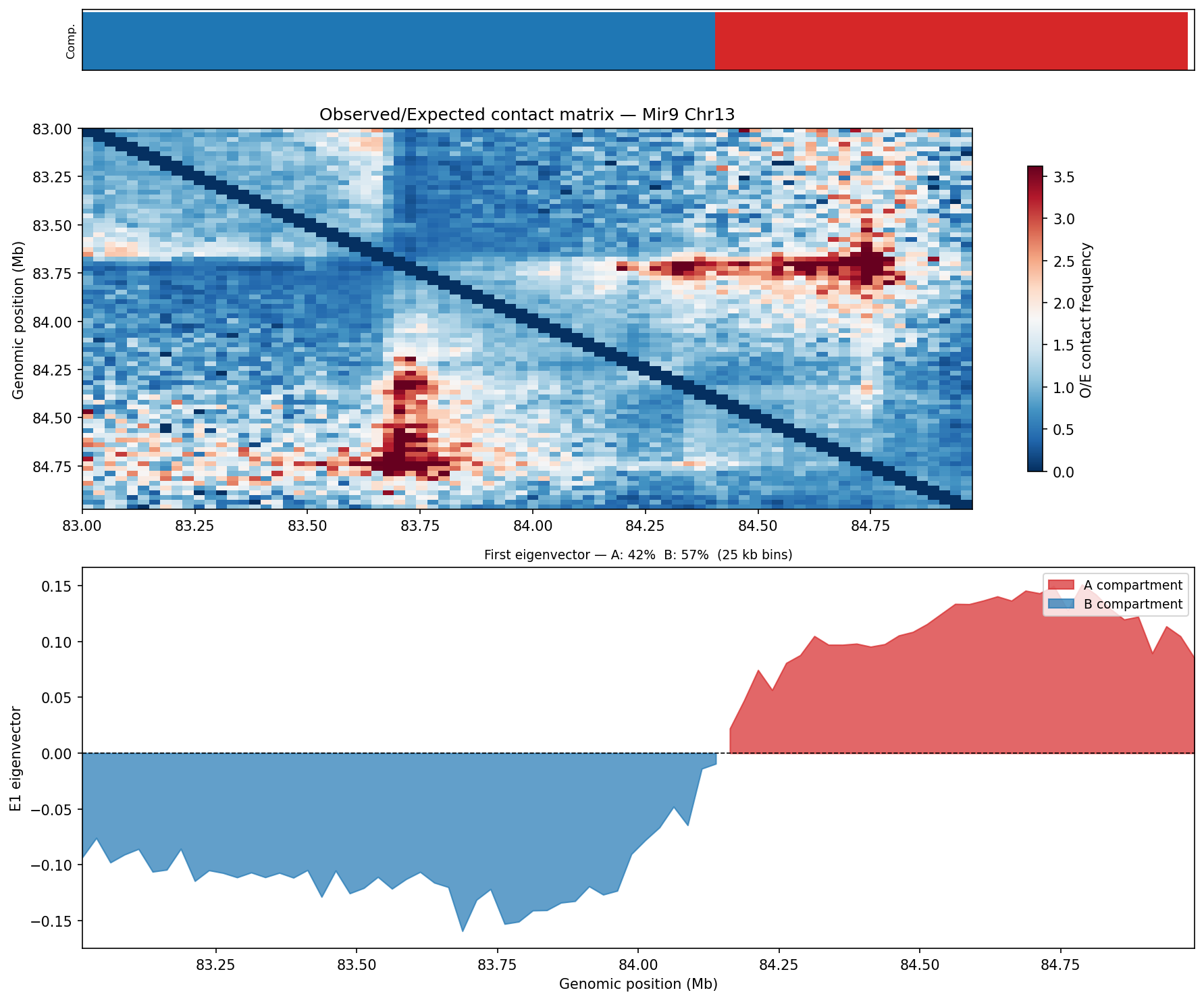
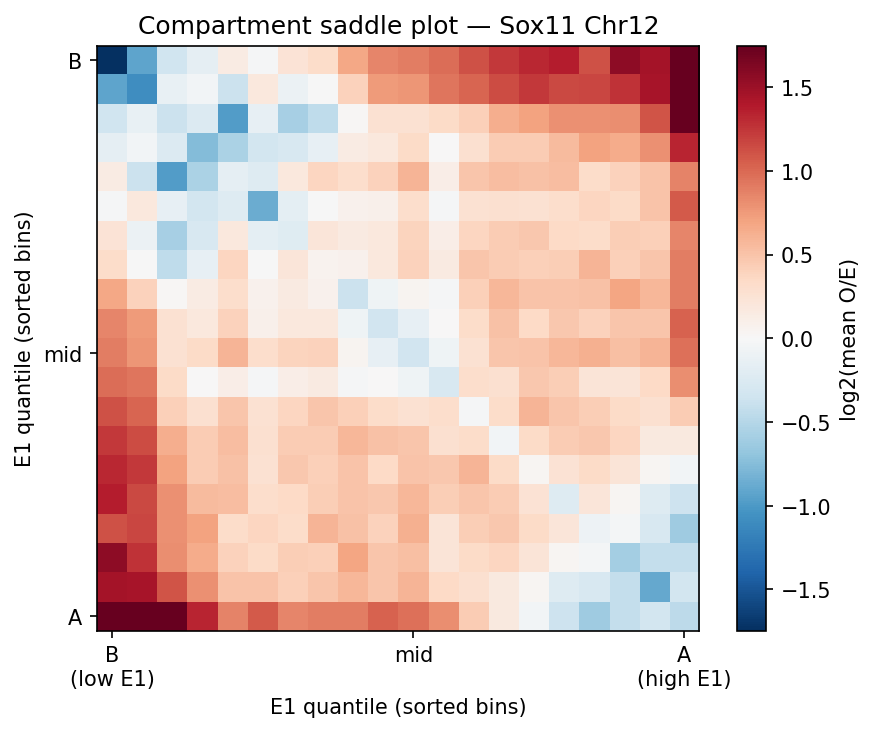
A/B Compartments (E1 Eigenvector)
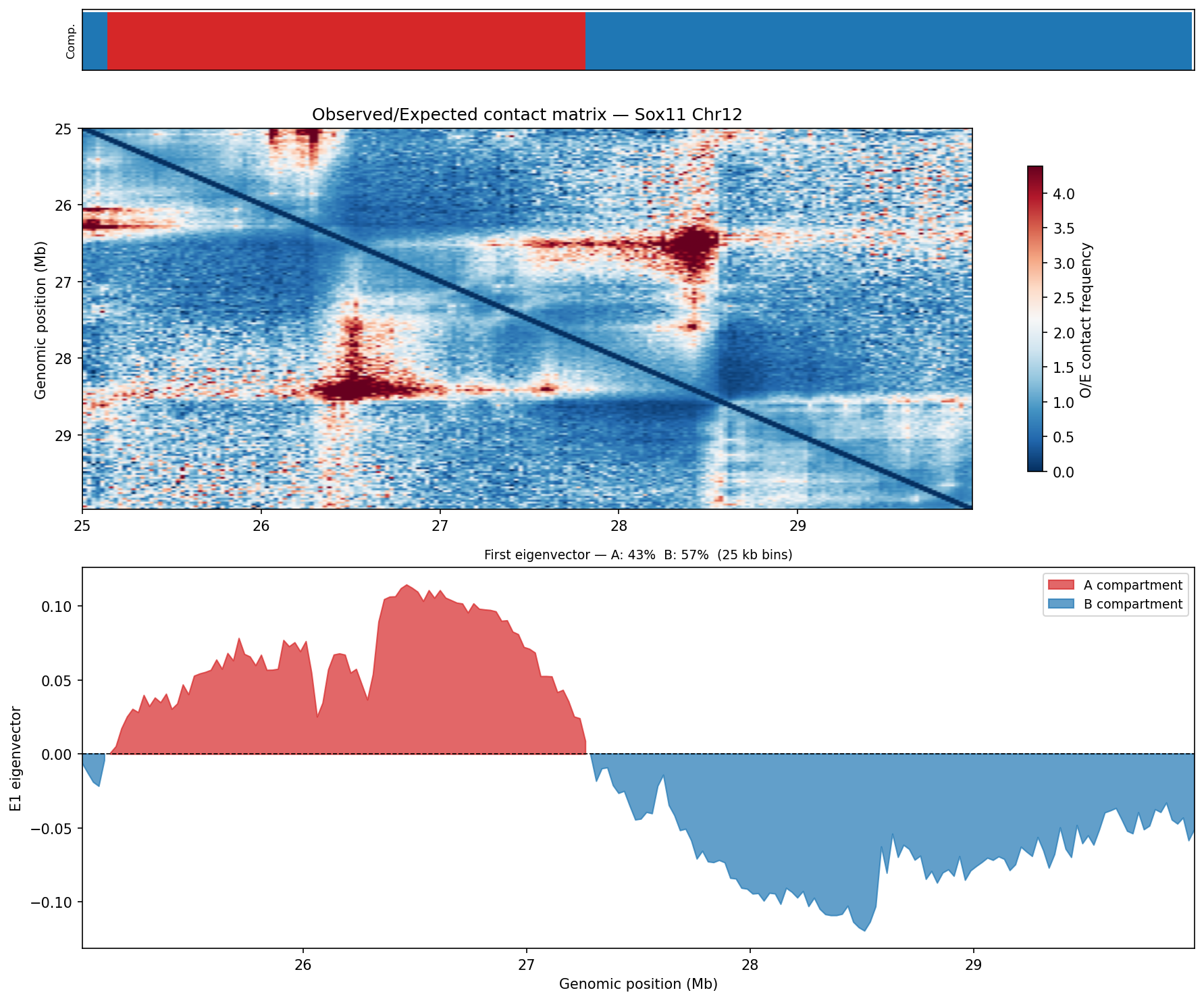
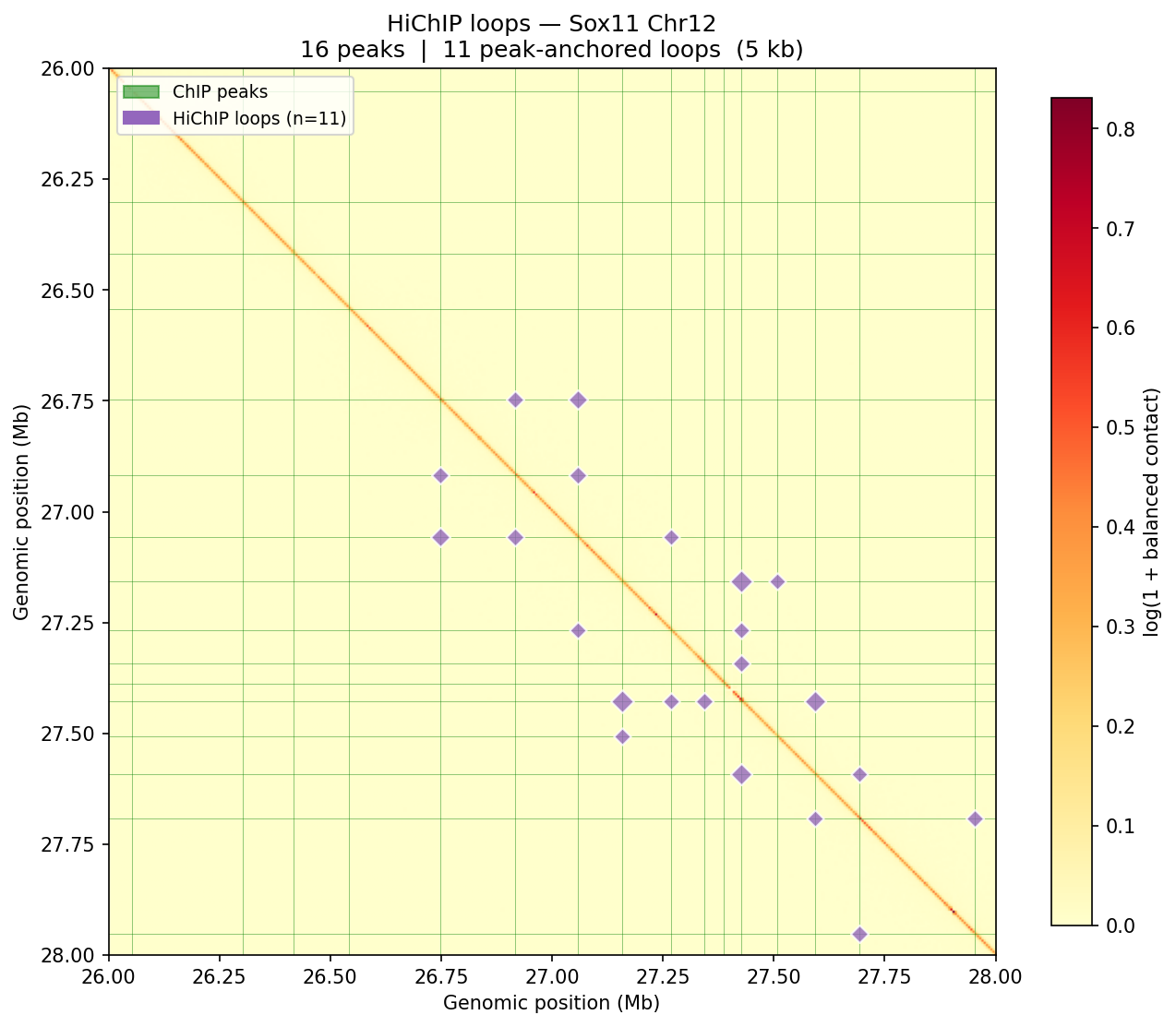
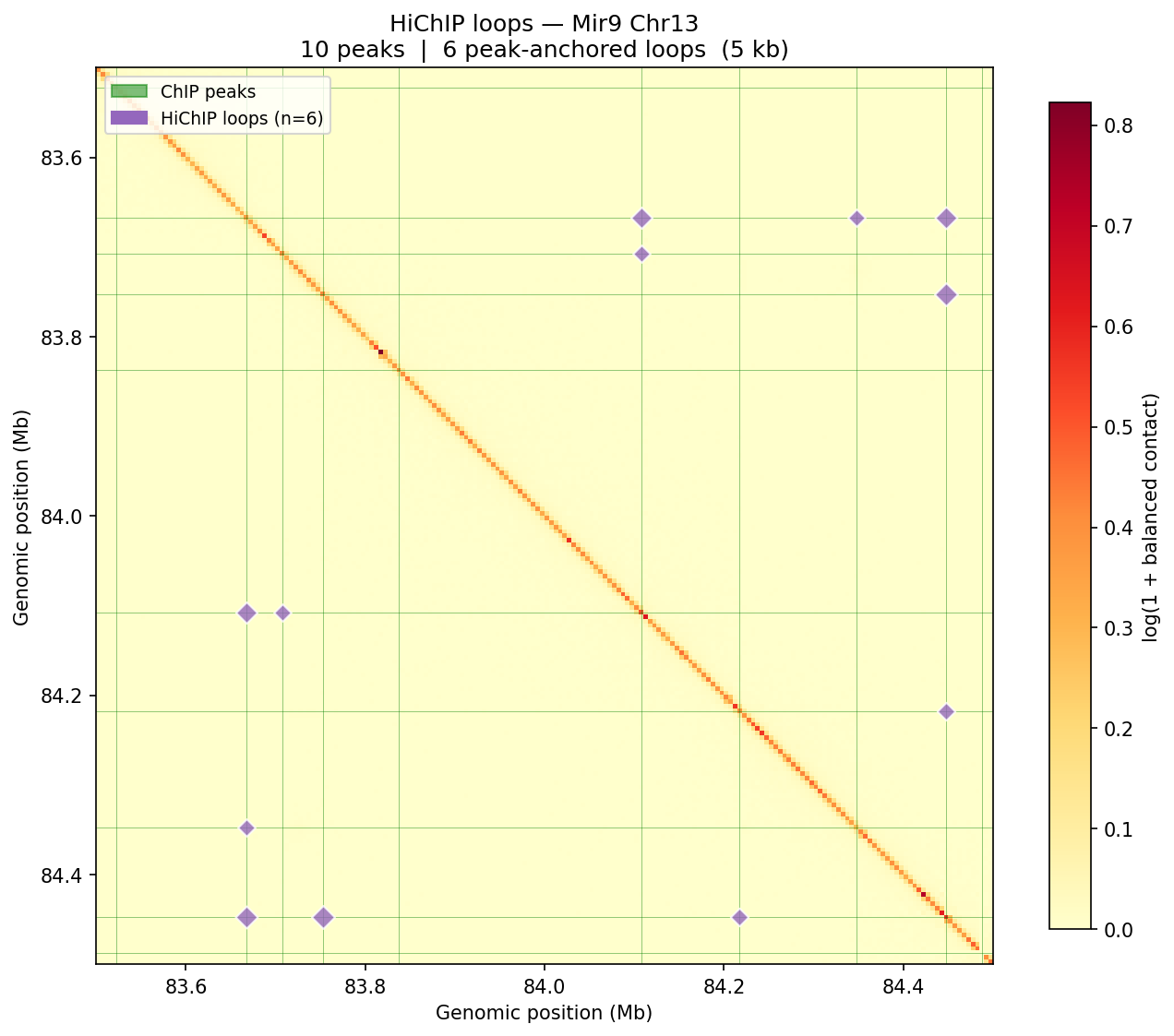
HiChIP — Peak-Anchored Loops
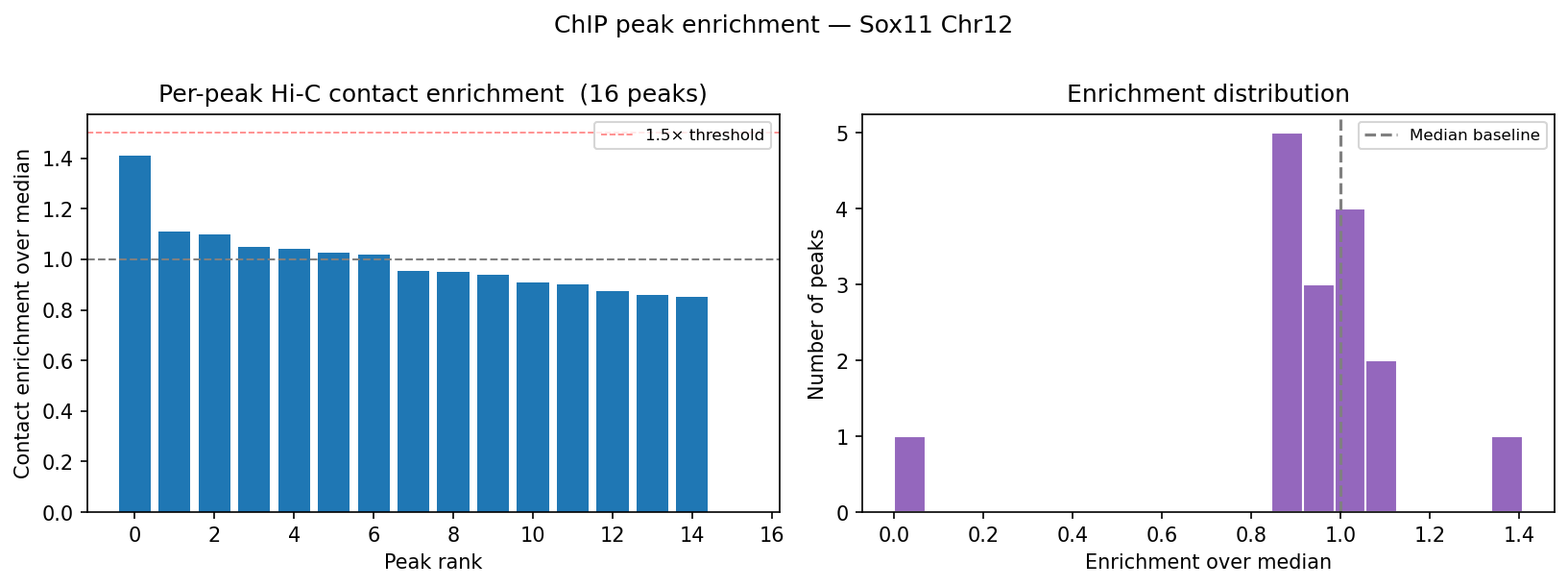
Capture Hi-C — CHiCAGO Model
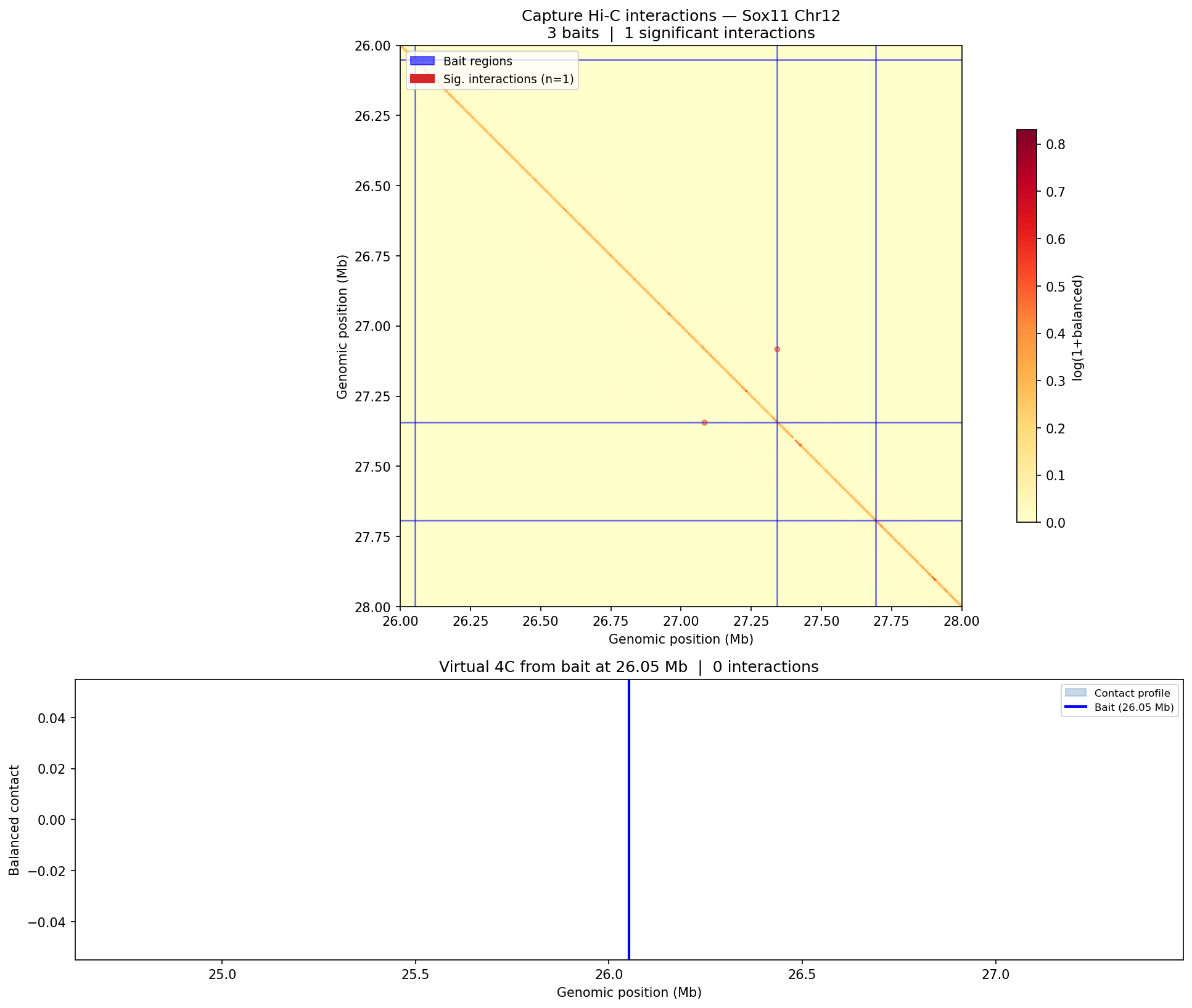
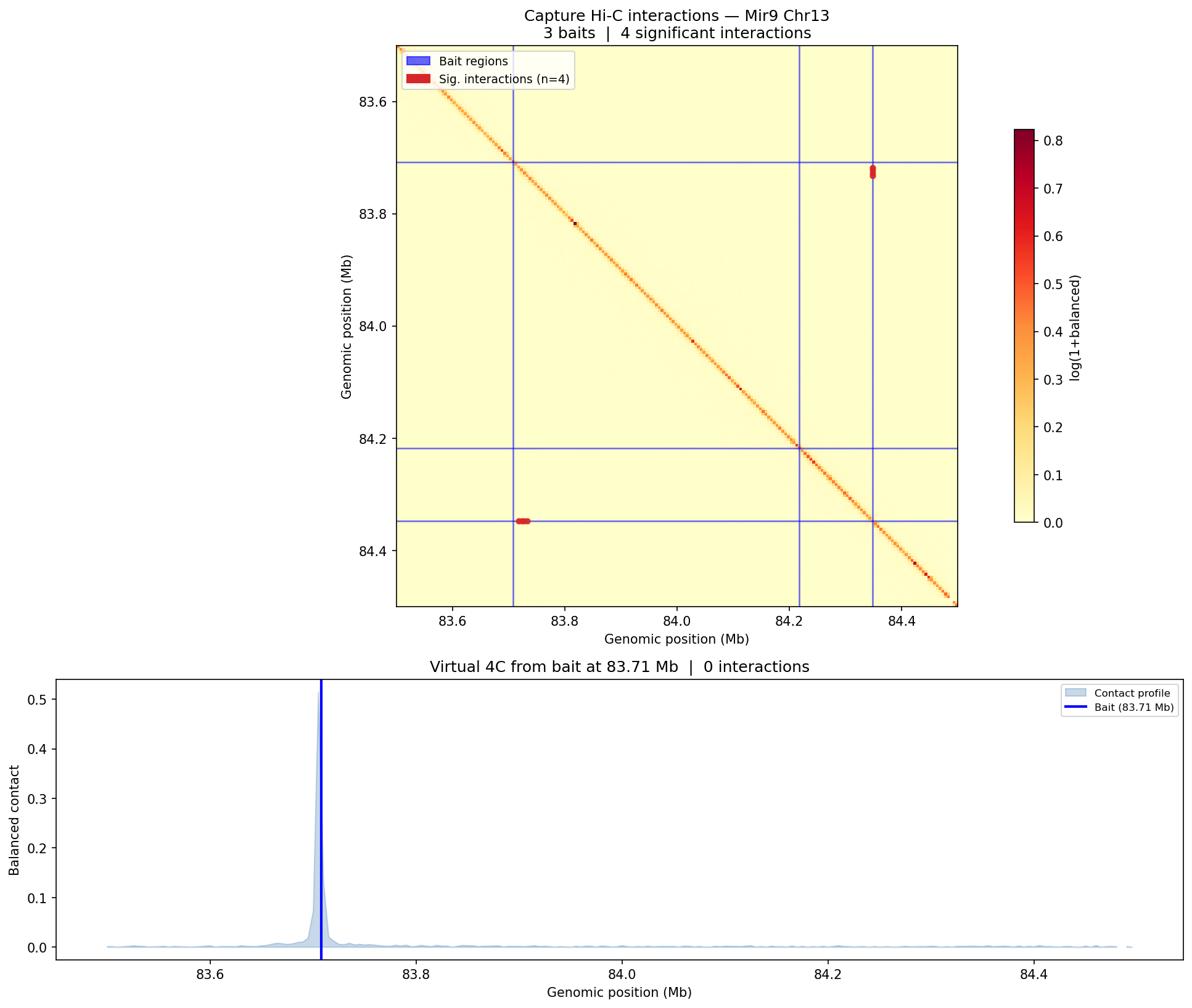
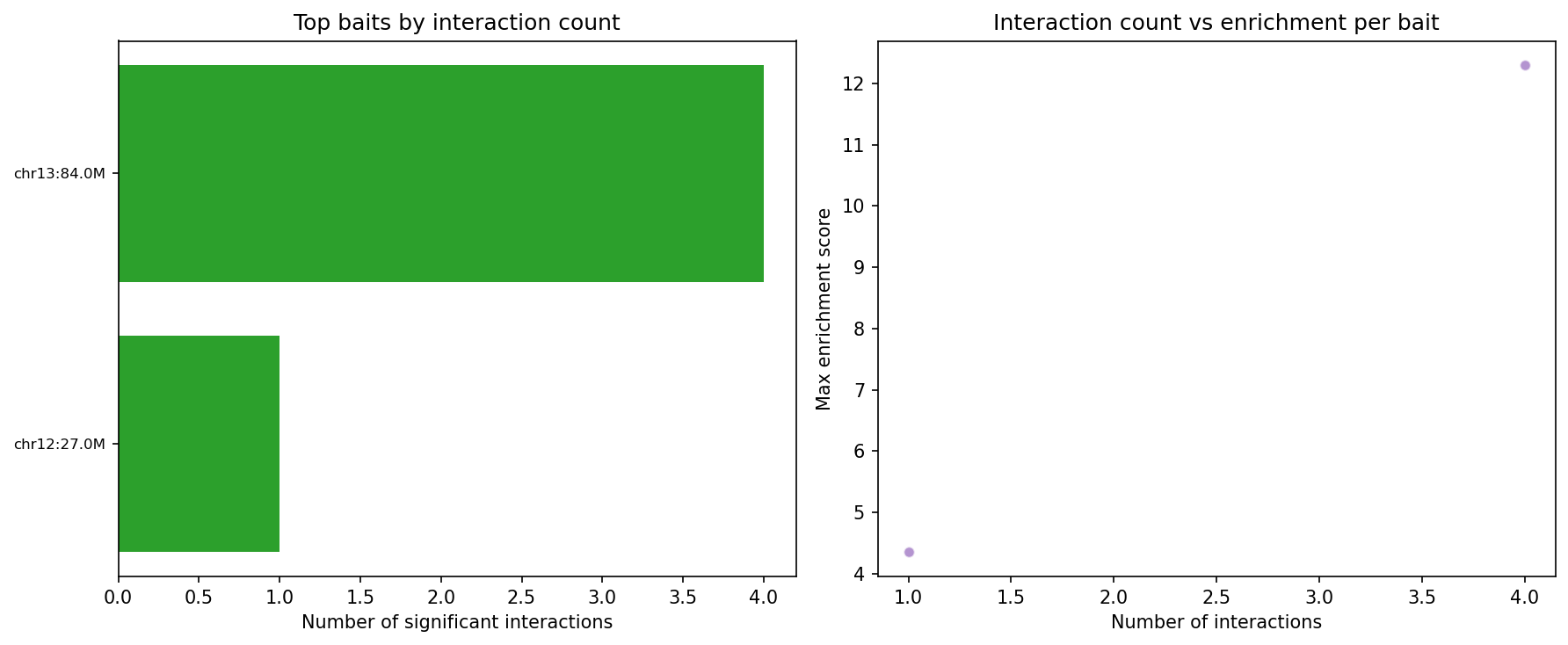
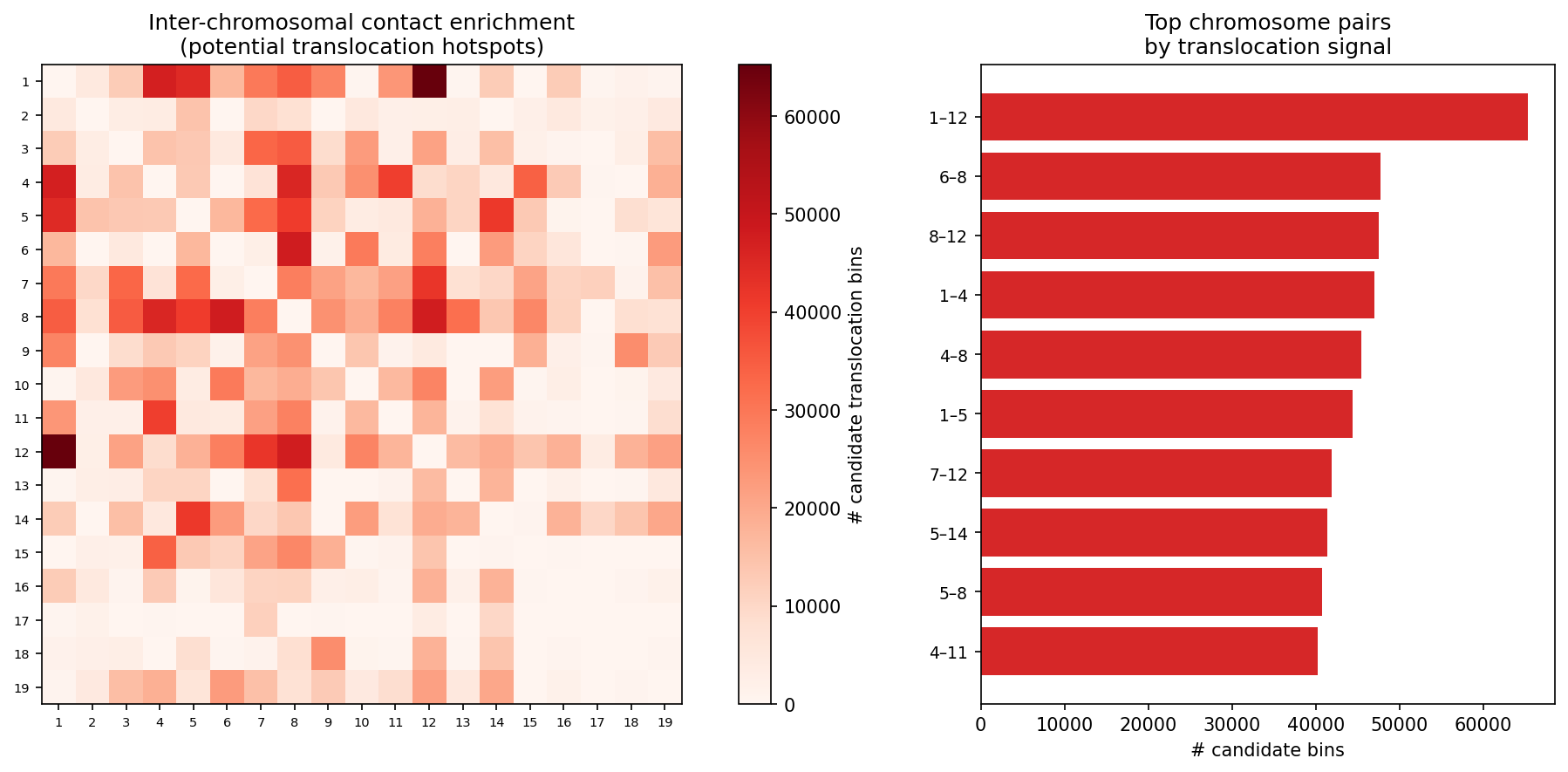
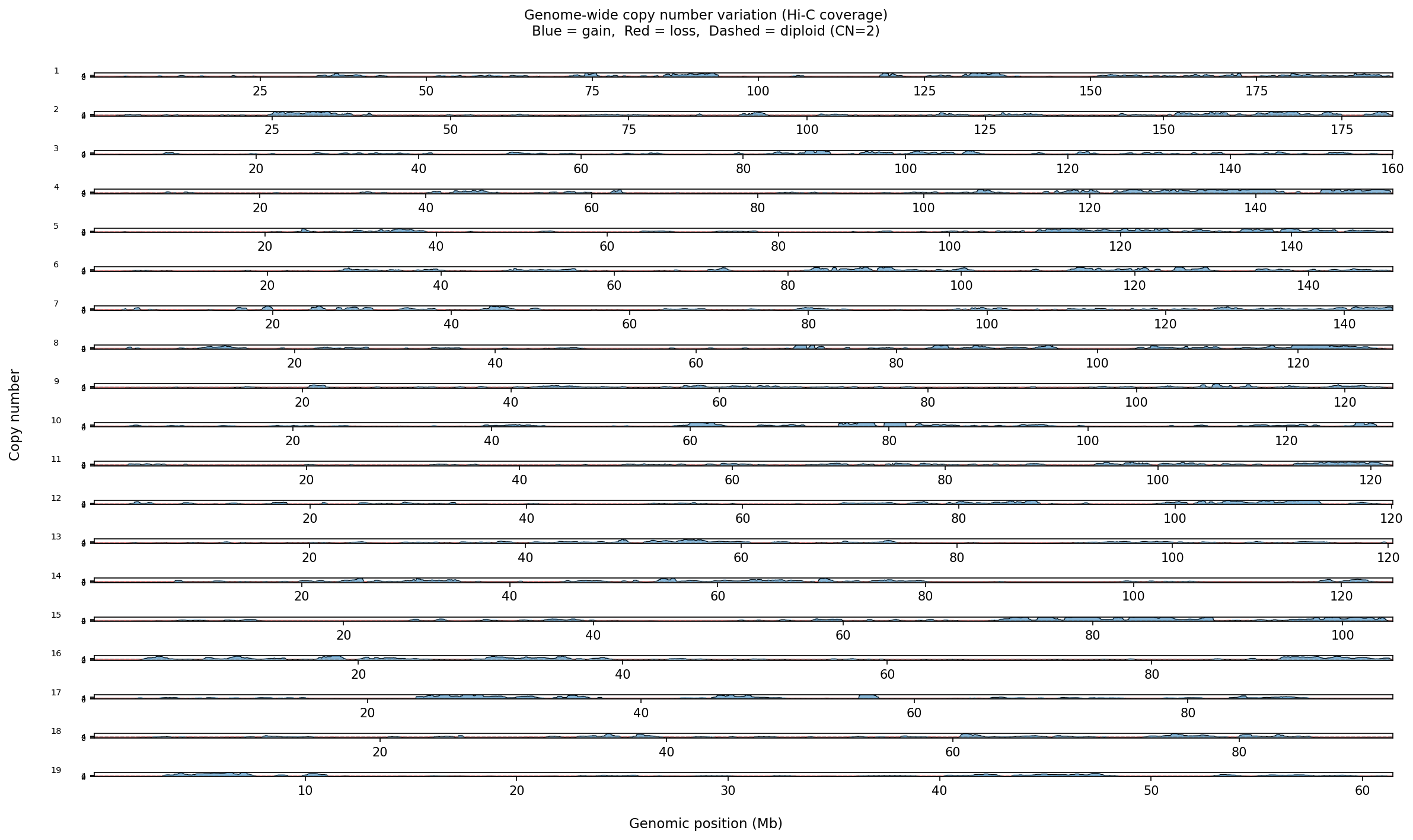
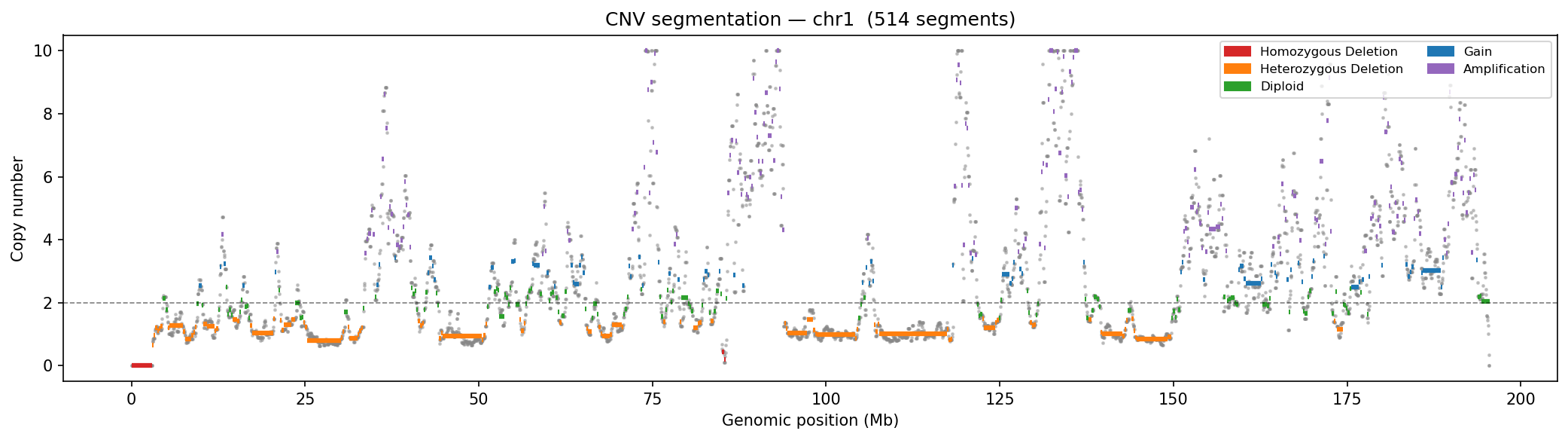
Structural Variants & CNV
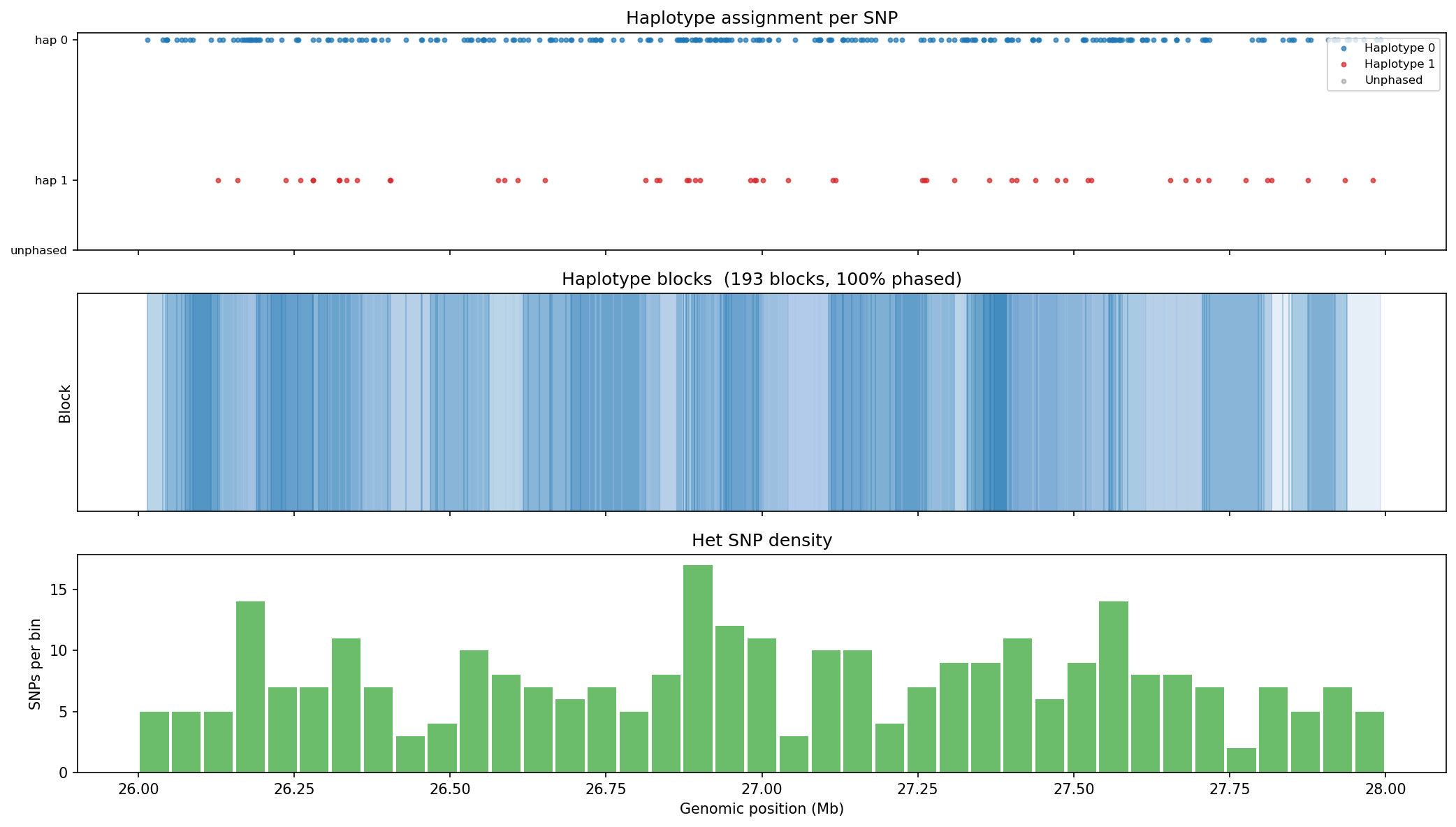
Haplotype Phasing & Variants
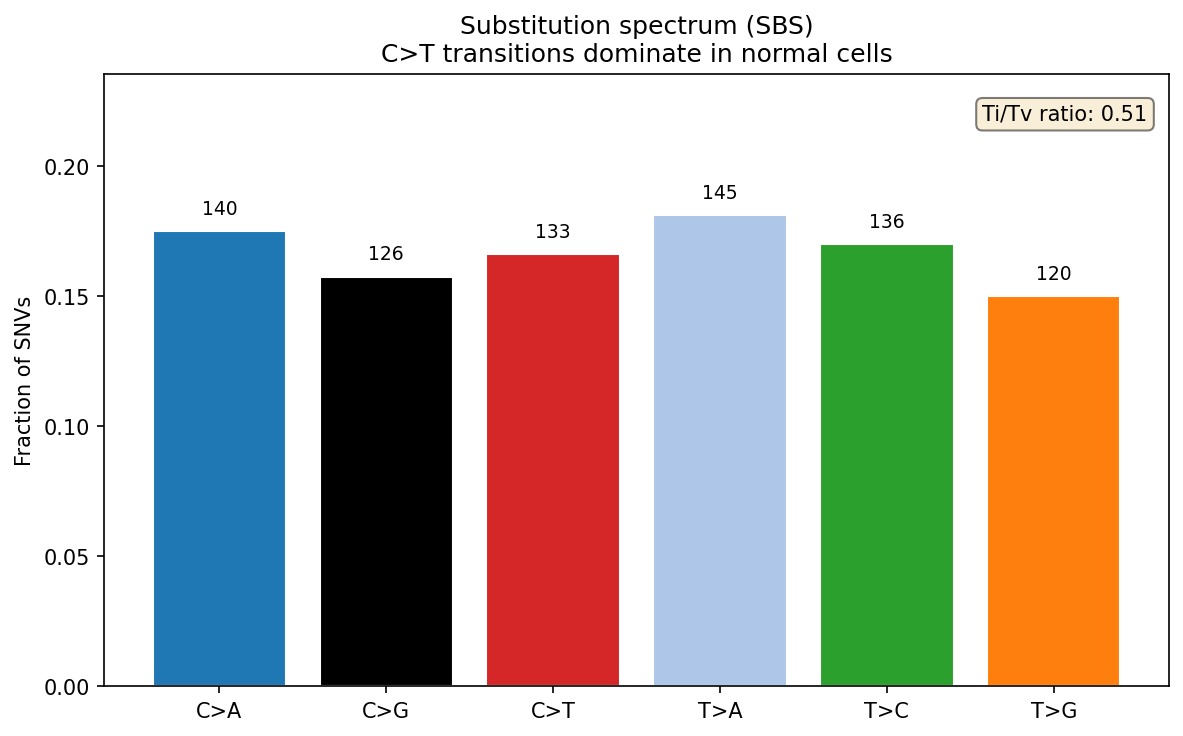
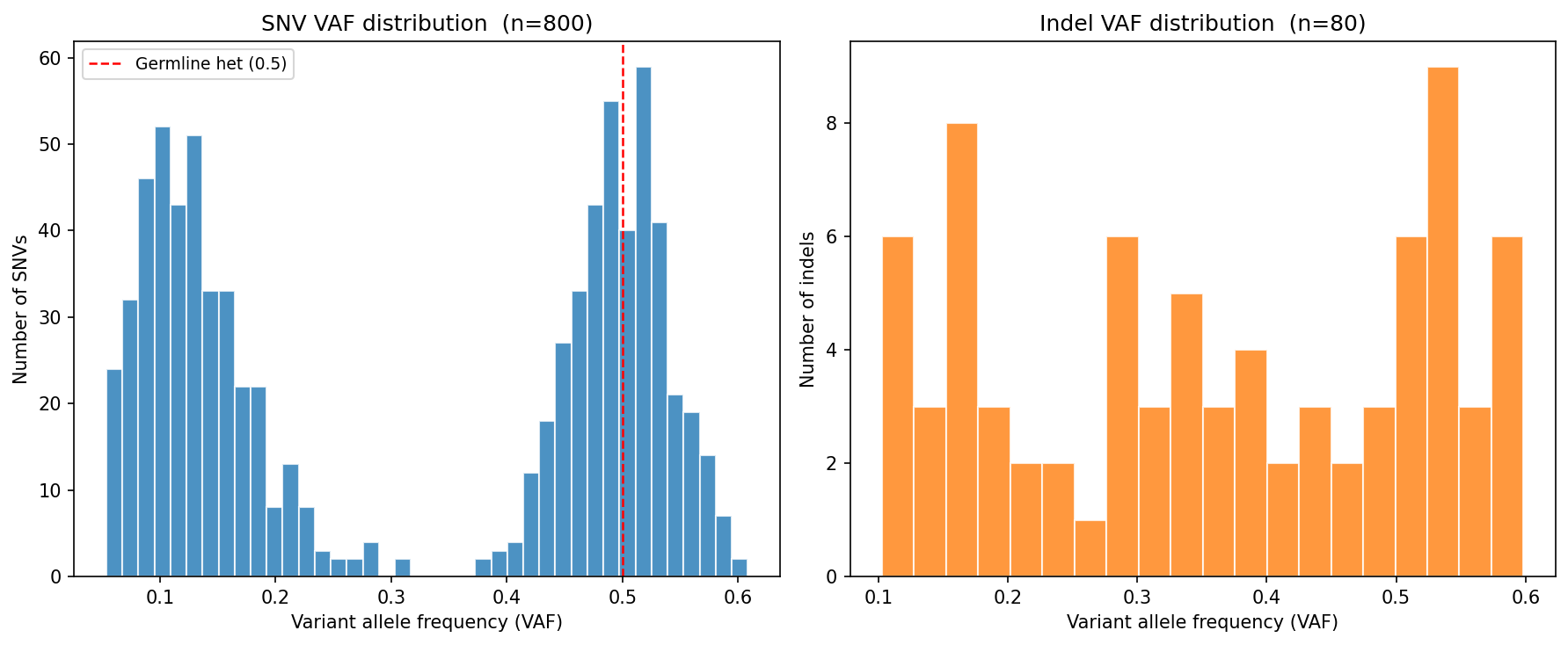
Mouse Insulator Deletion Analysis UPDATED
In silico prediction of insulator deletion effects in mouse (mm10) across two cell types, run at the suggestion of the PI. Regions provided by collaborators studying inner ear development.
Cell types:
CL:0000207 olfactory receptor cell (primary sensory neuron, closest proxy to inner ear hair cells) &
EFO:0004038 mouse embryonic stem cell (PI-suggested baseline).
Note: UBERON:0001846 (internal ear) exists in AlphaGenome for CAGE only — not contact maps. No otic placode GO/UBERON terms are in the mouse contact map training data (only 8 mouse tracks total).
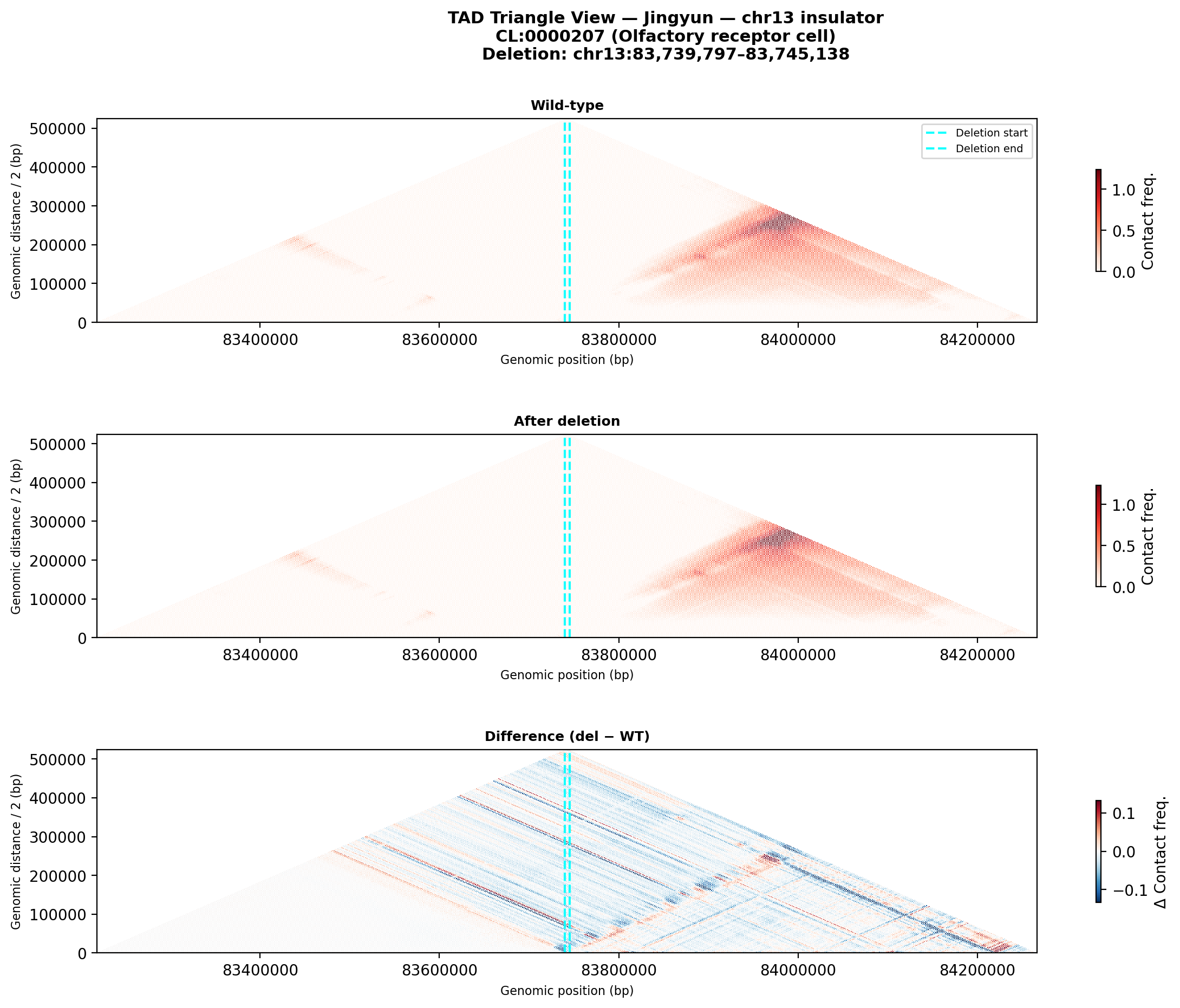
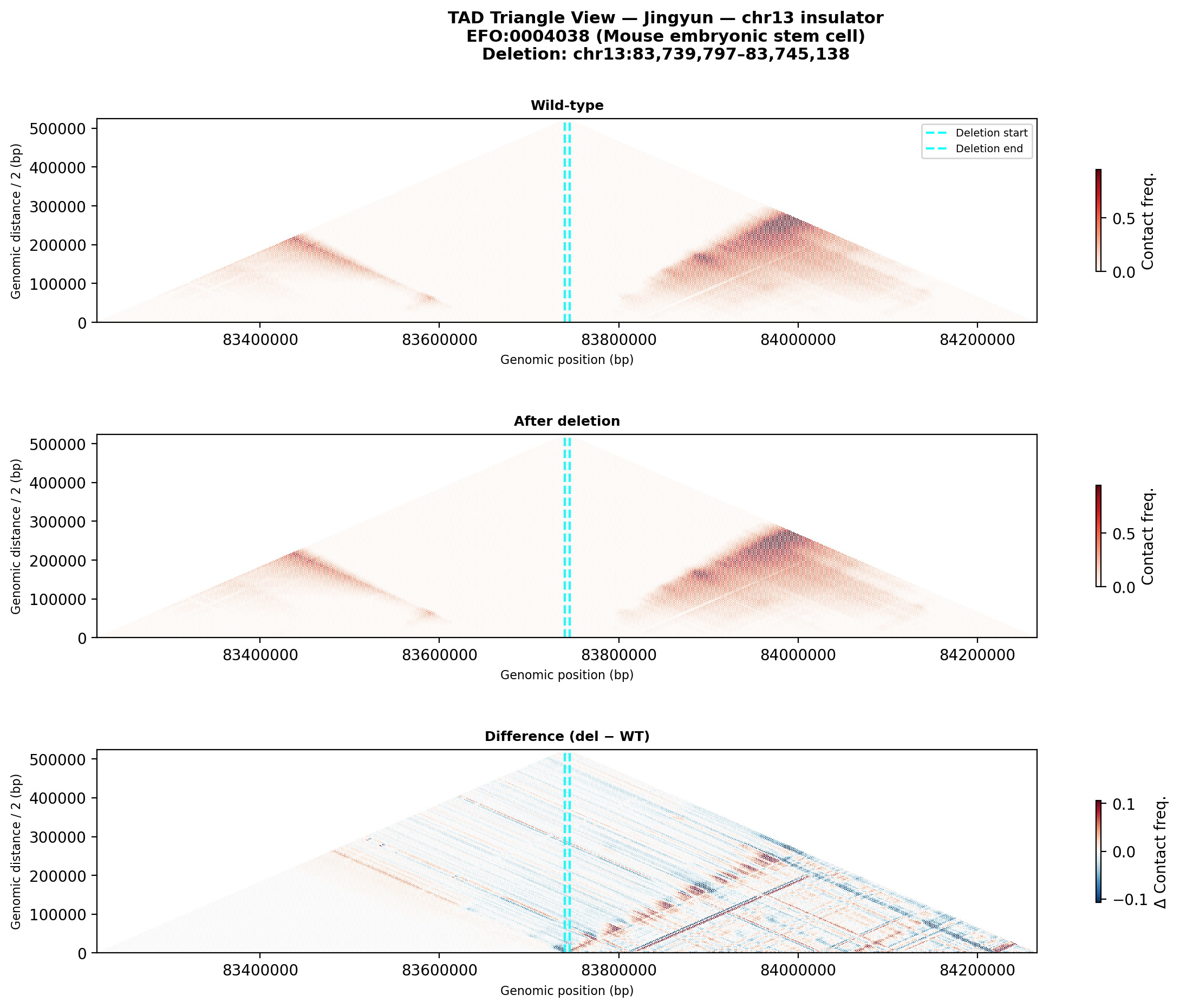
Jingyun — chr13:83,739,797–83,745,138 (confirmed insulator, 5,342 bp)
Coordinates confirmed directly by Jingyun. Full 2-TAD span: chr13:81,760,002–85,200,000 (3.4 Mb, larger than the AlphaGenome 1 Mb API limit — predictions shown for a 1 Mb window centred on the deletion site).
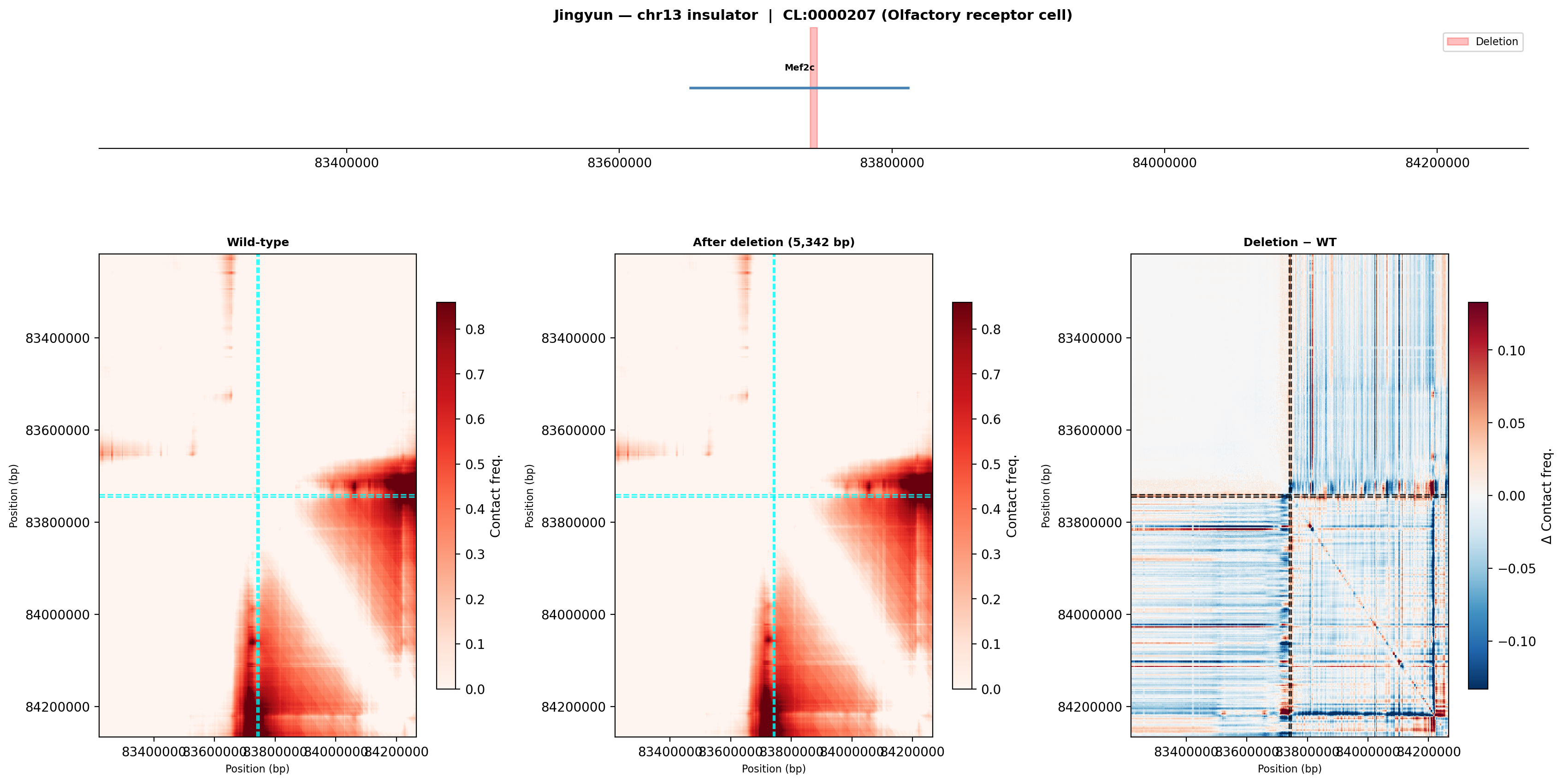
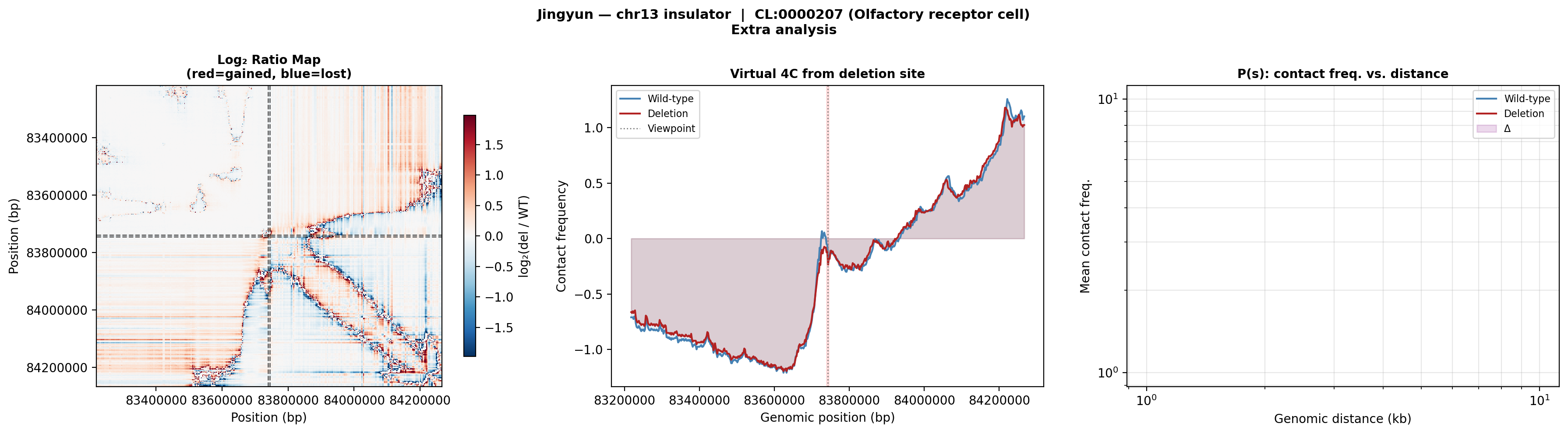
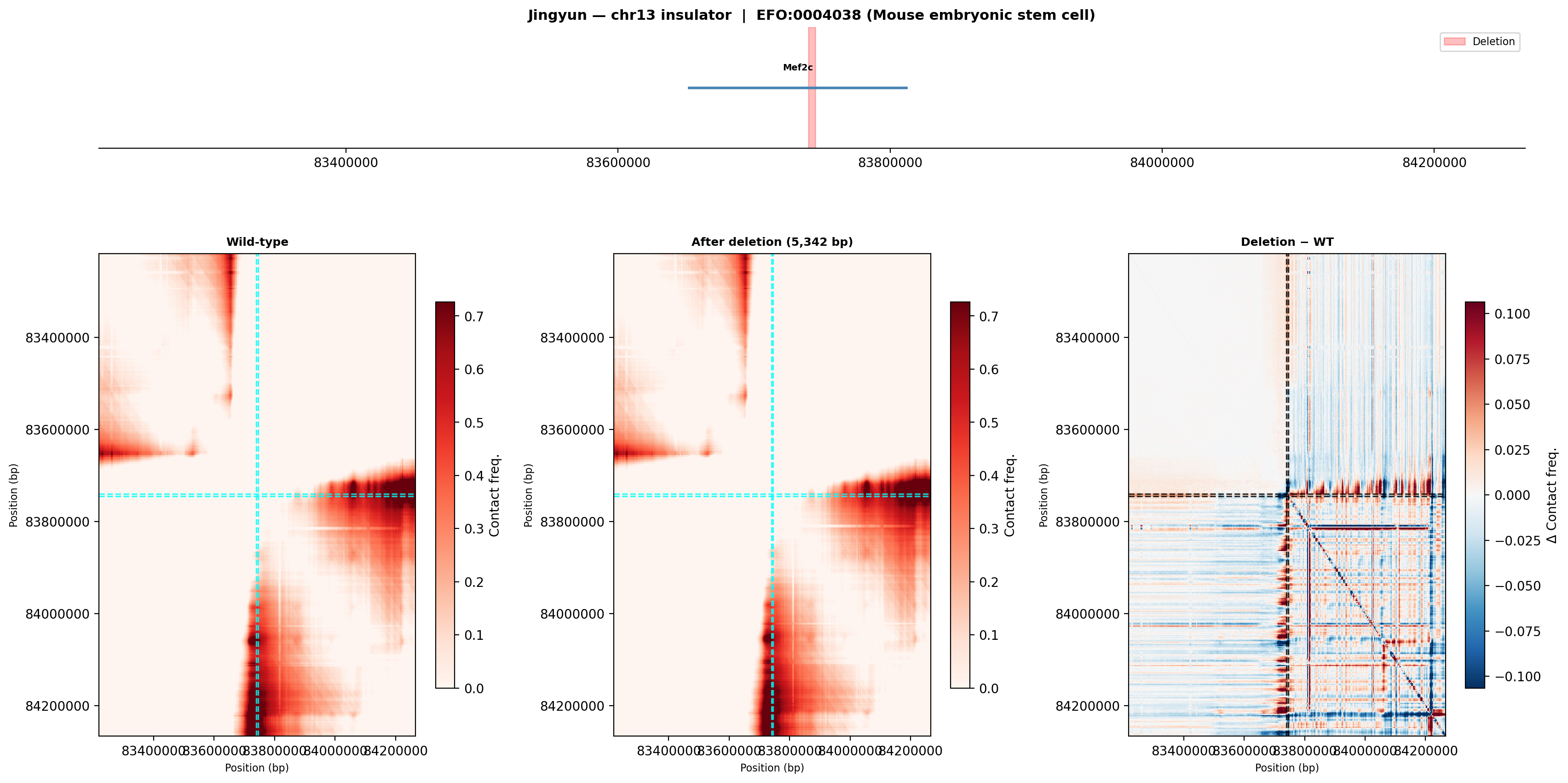
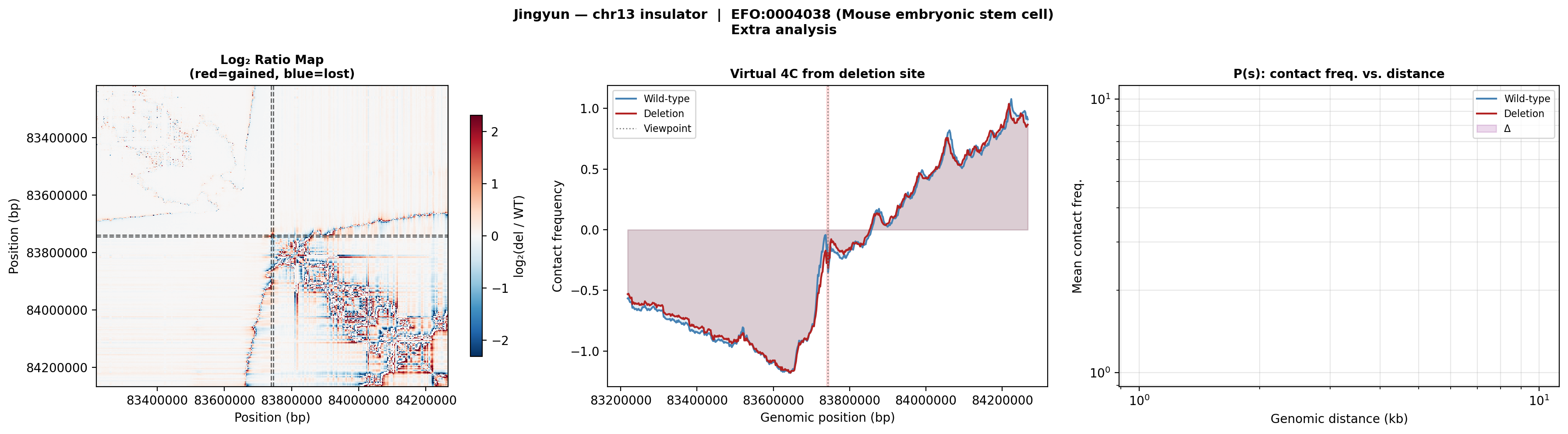
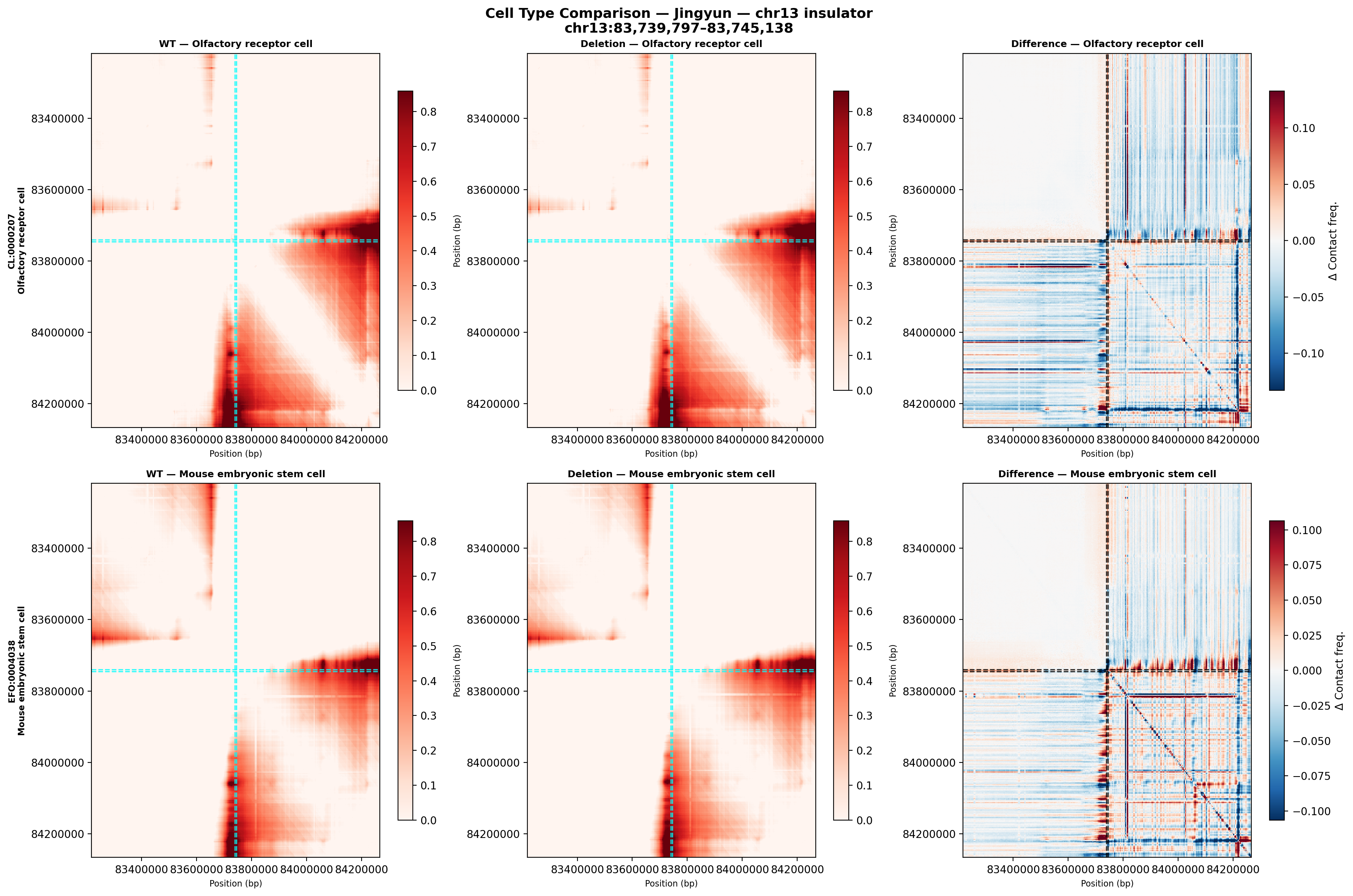
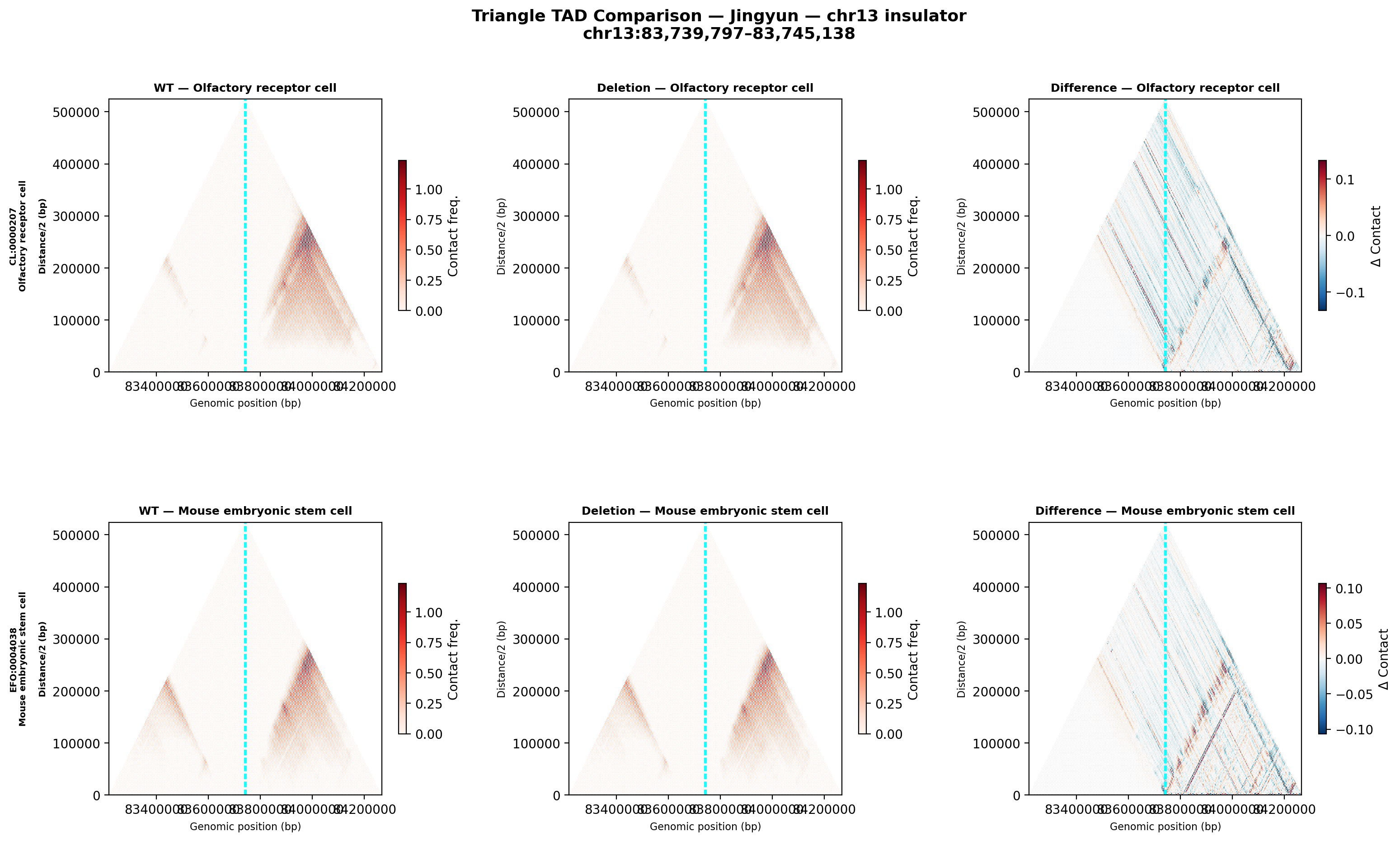
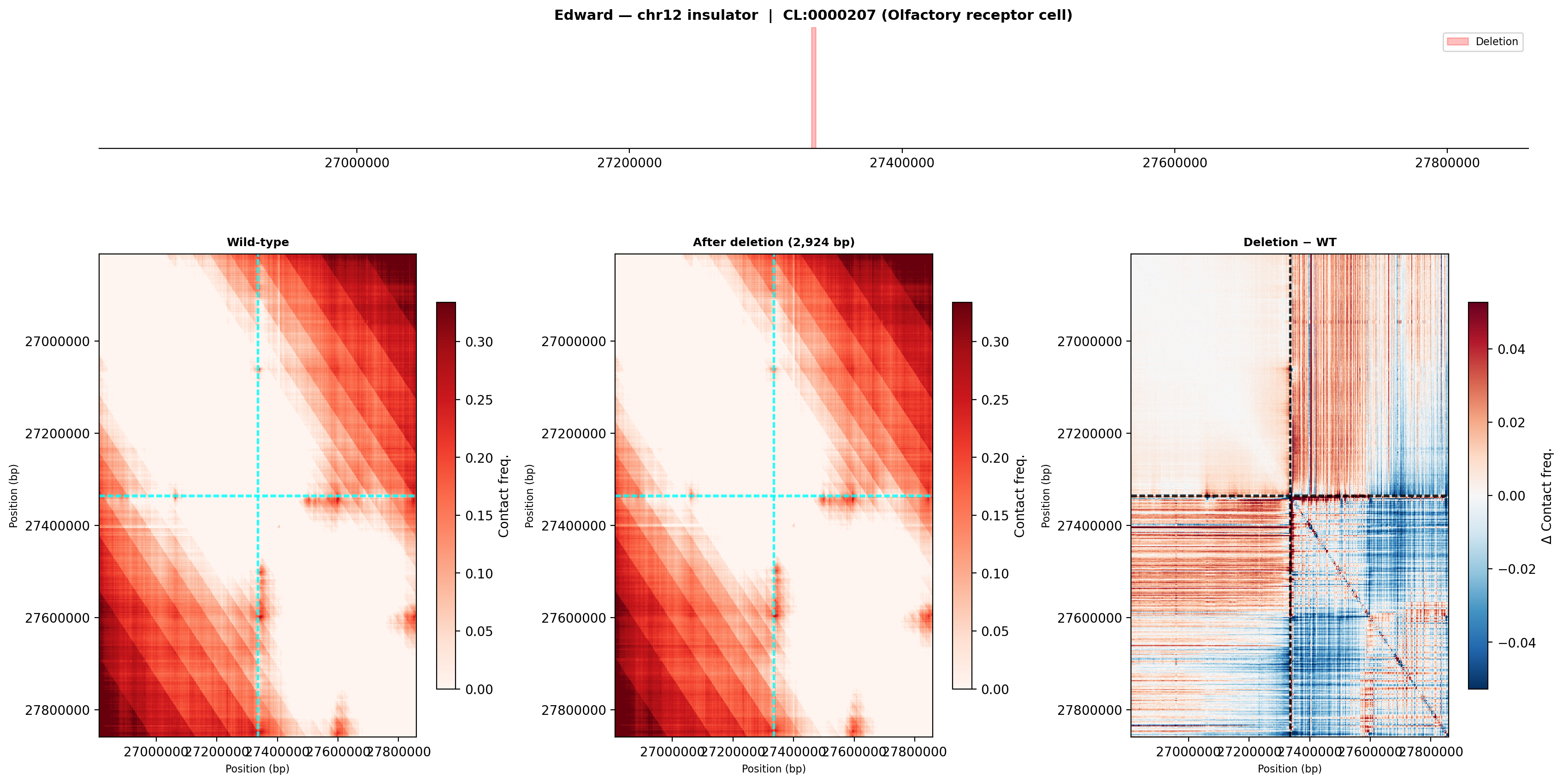
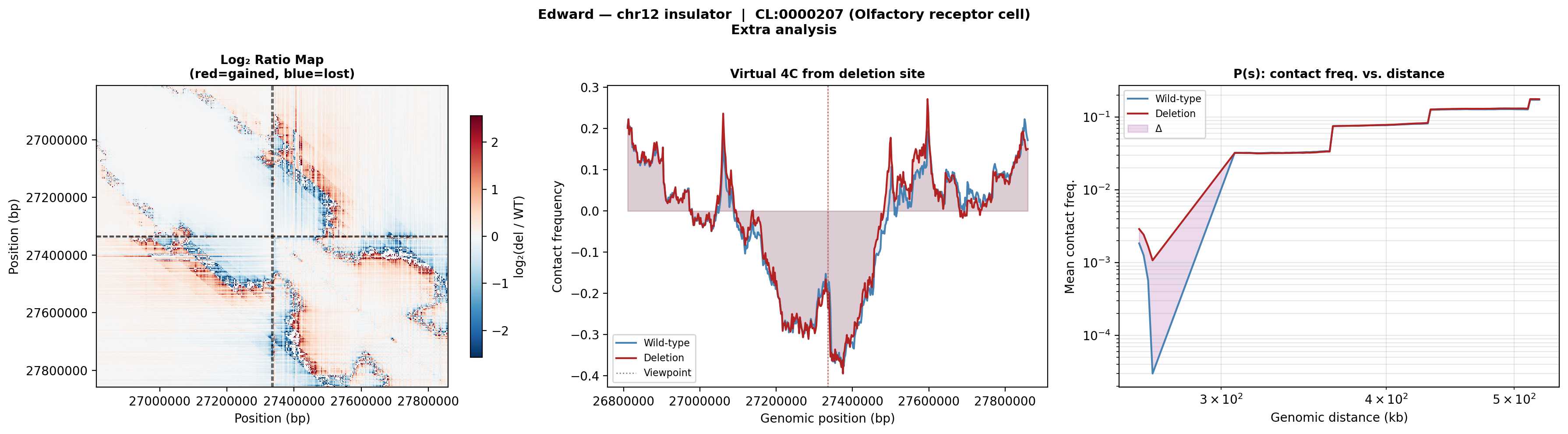
Edward — chr12:27,333,532-27,336,455 (intergenic, 2,924 bp)
No protein-coding genes annotated in the 1 MB window (GENCODE M23) — deletion targets an intergenic regulatory element or insulator.
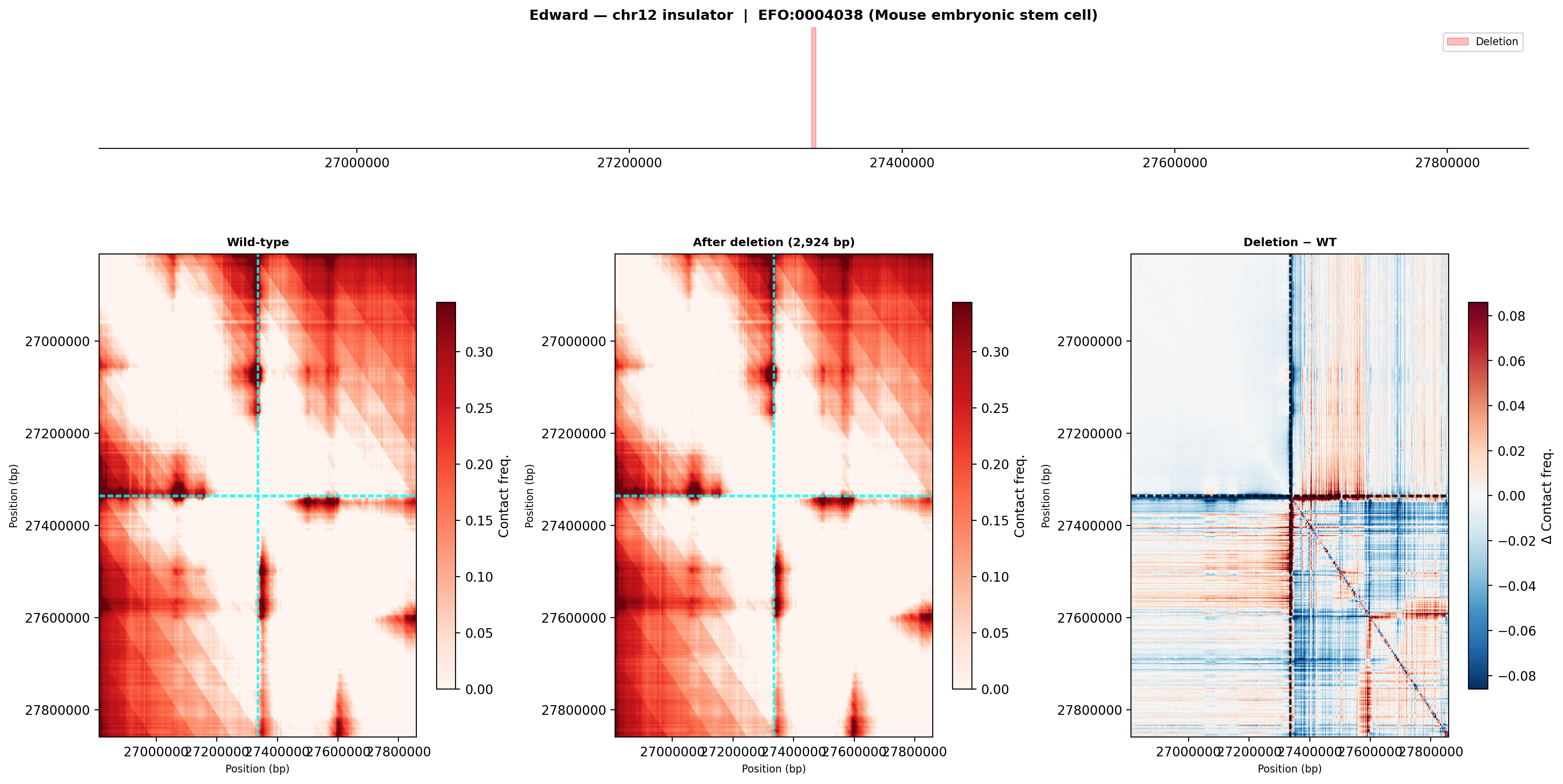
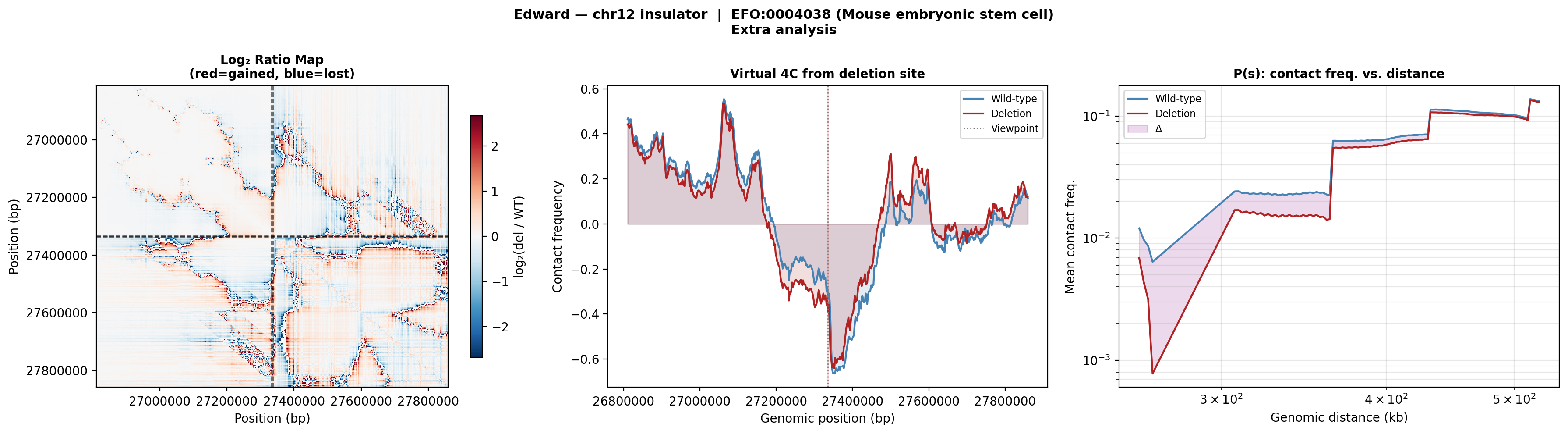
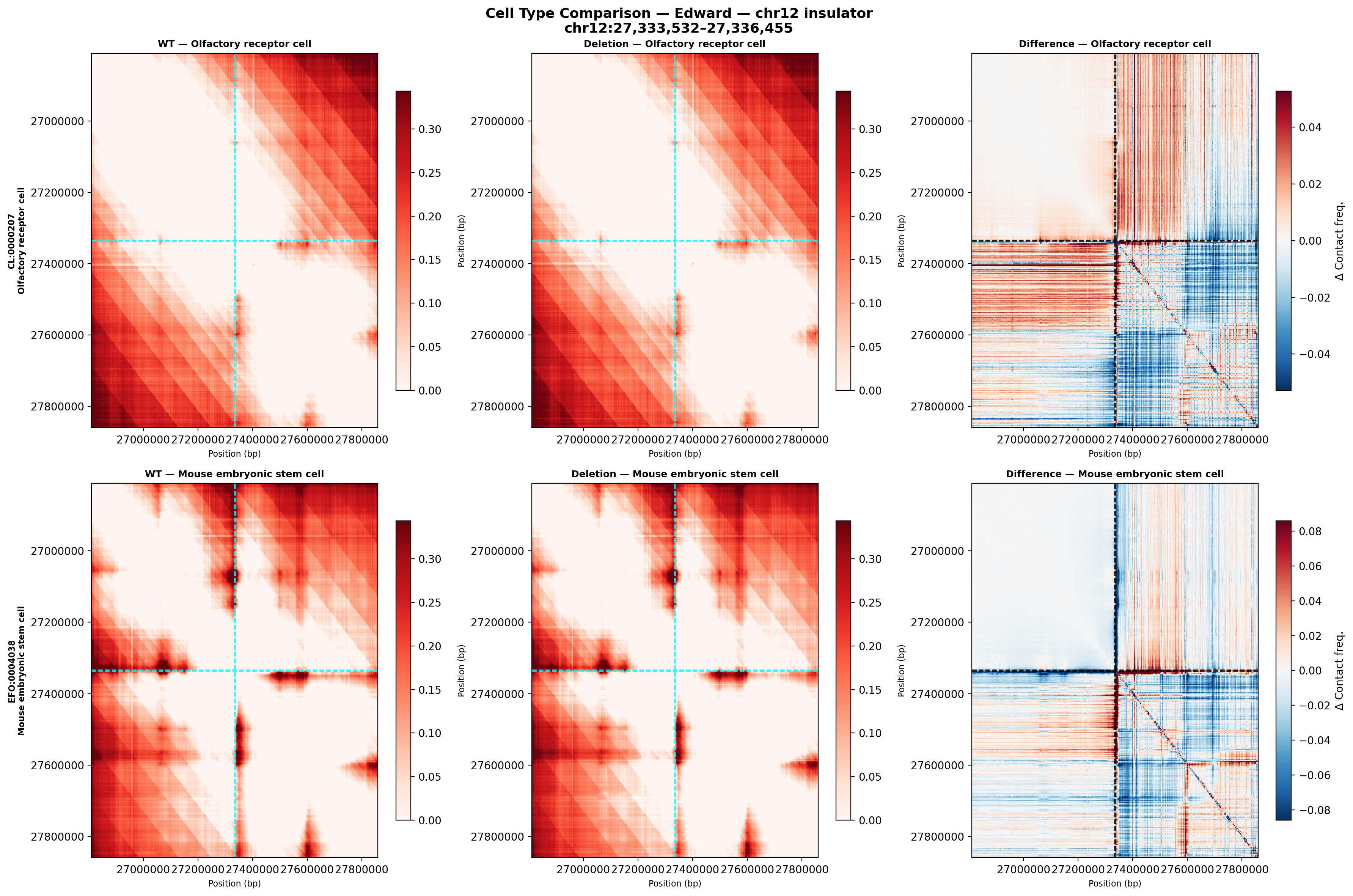
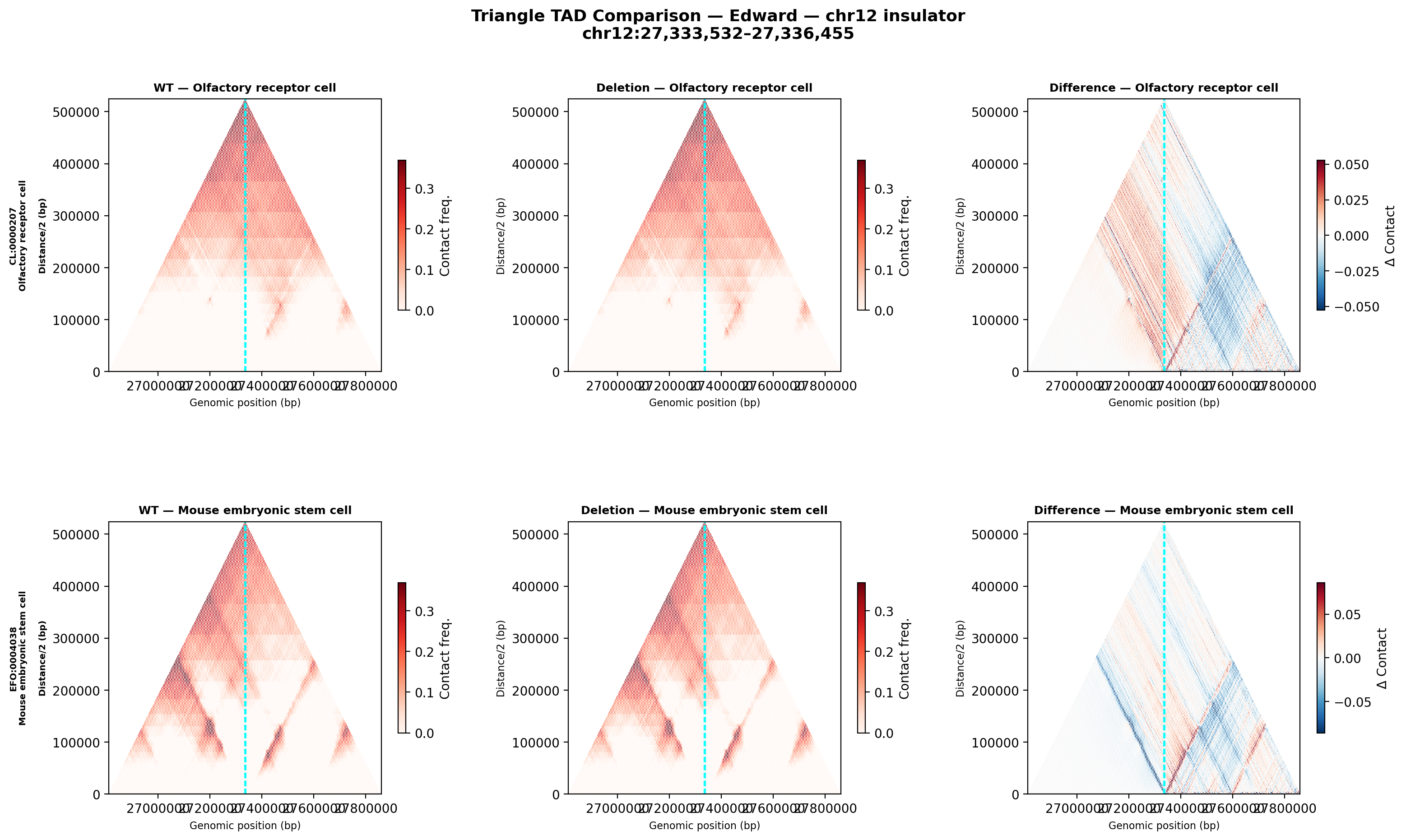
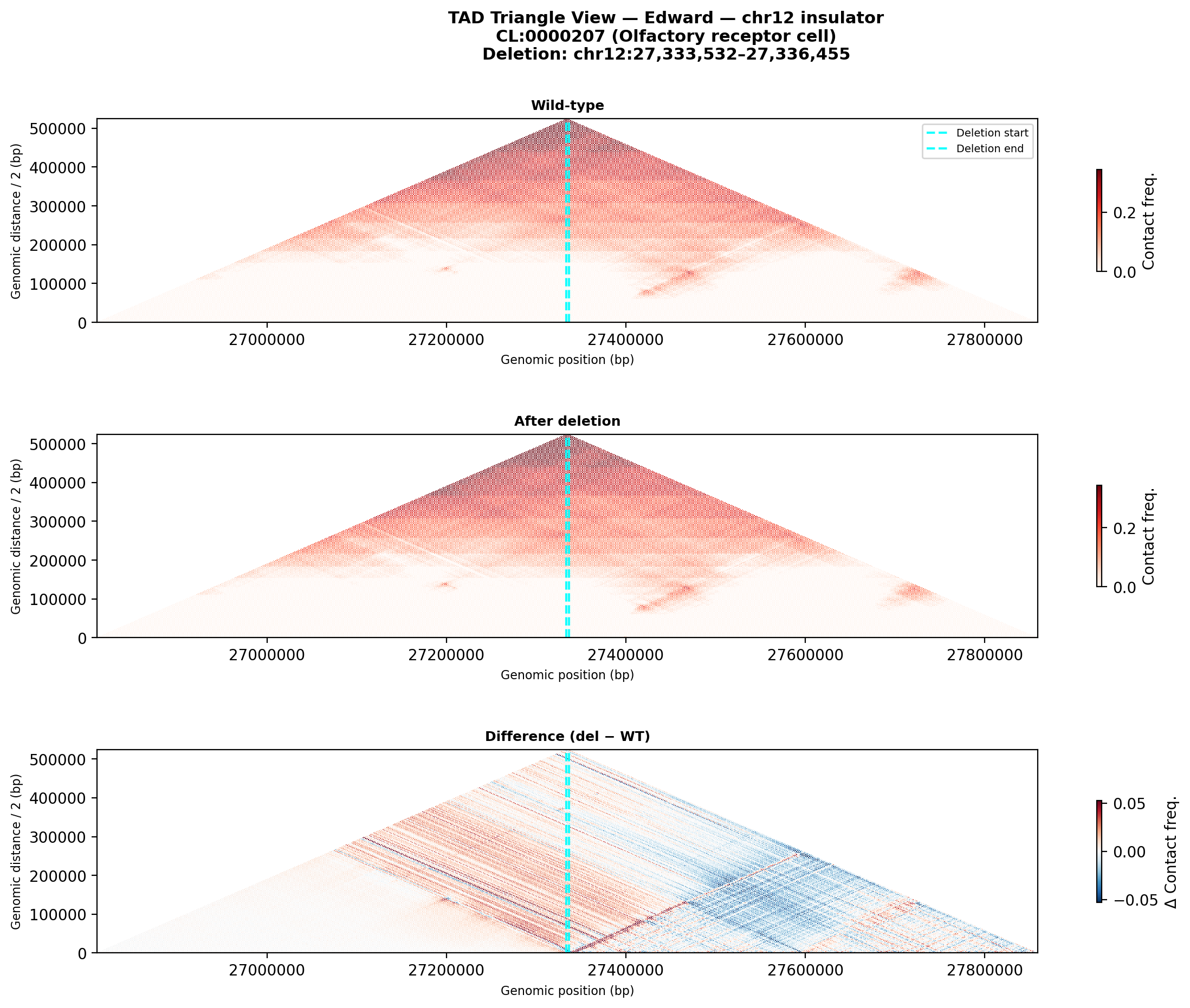
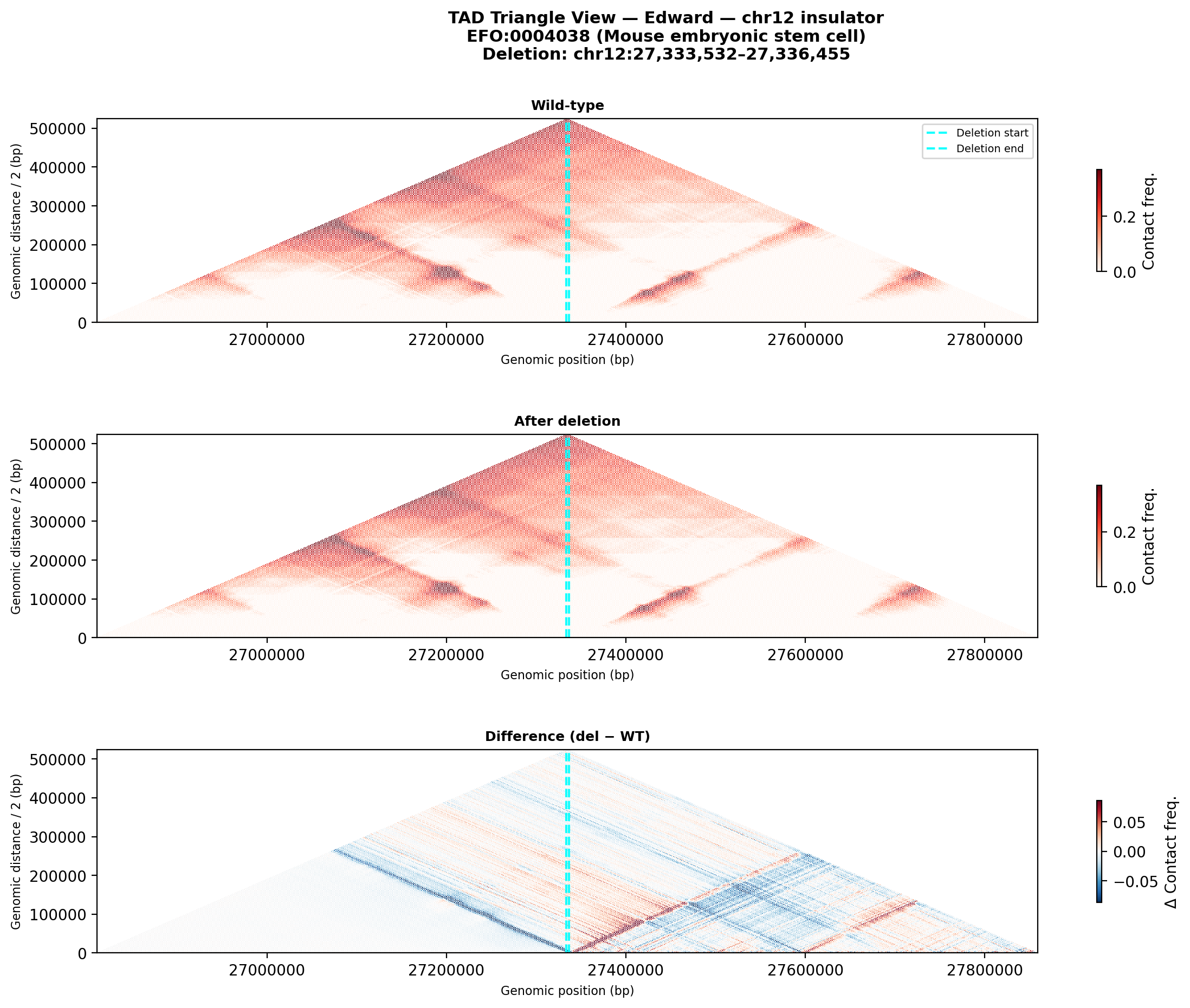
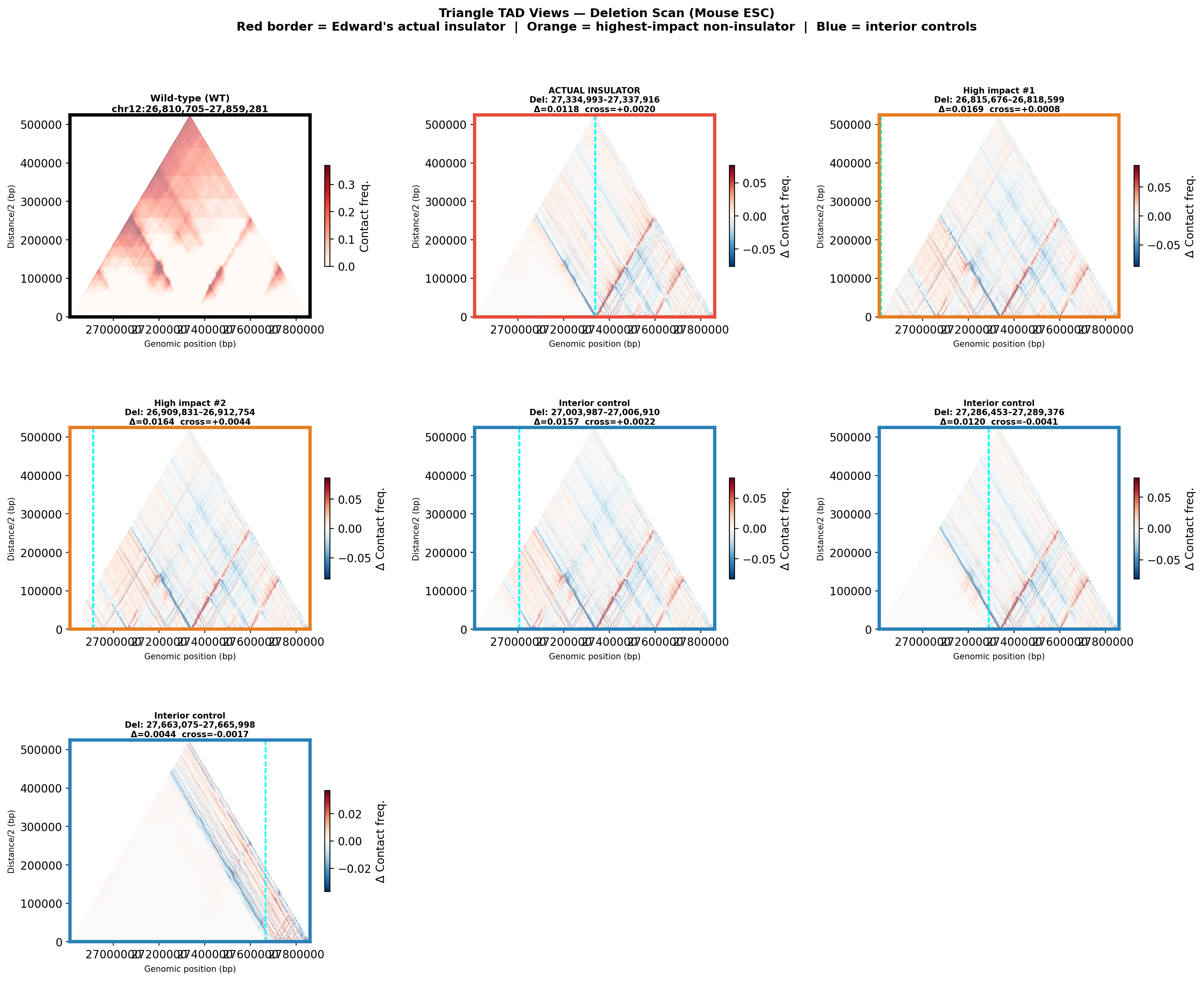
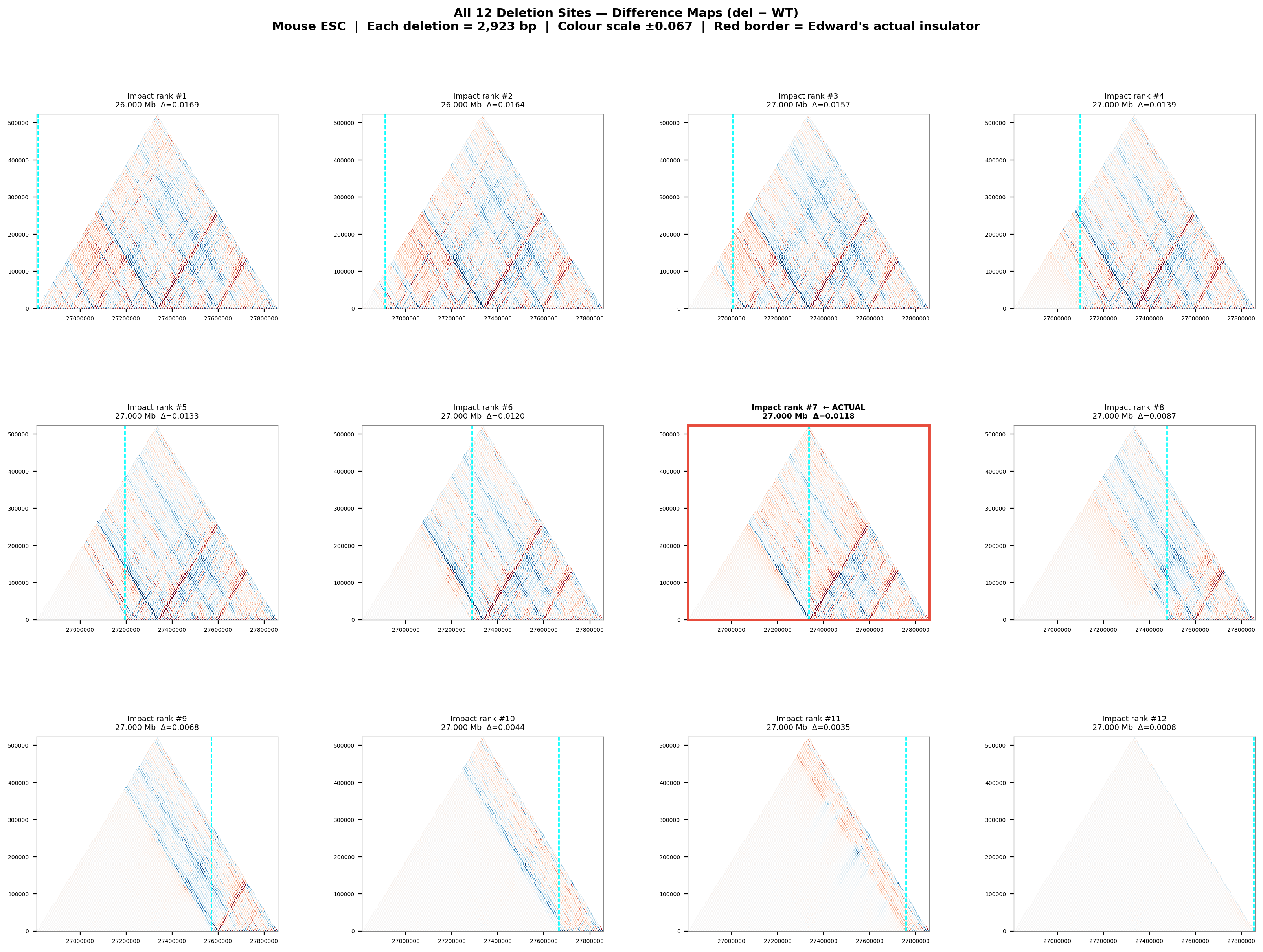
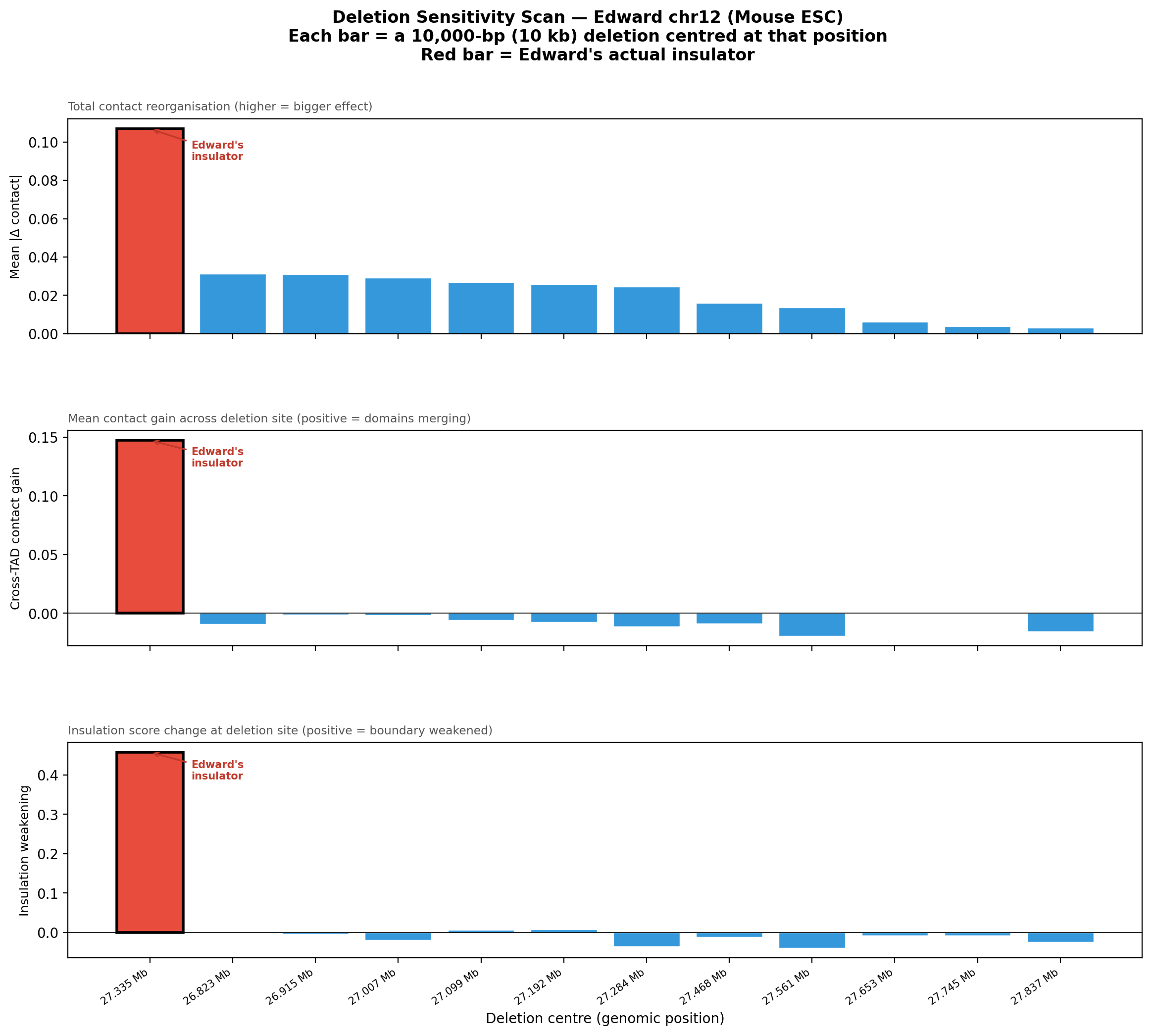
Deletion Sensitivity Scan — Edward chr12 NEW
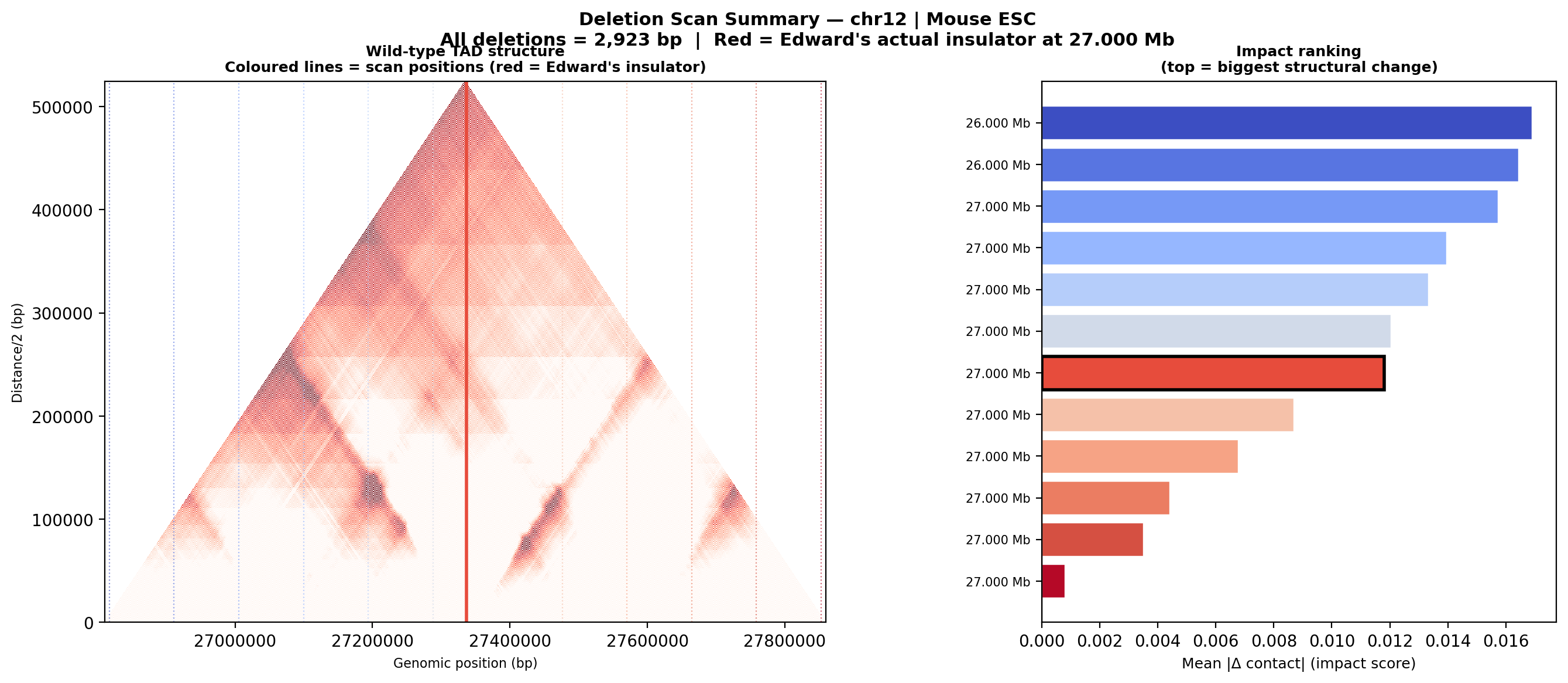
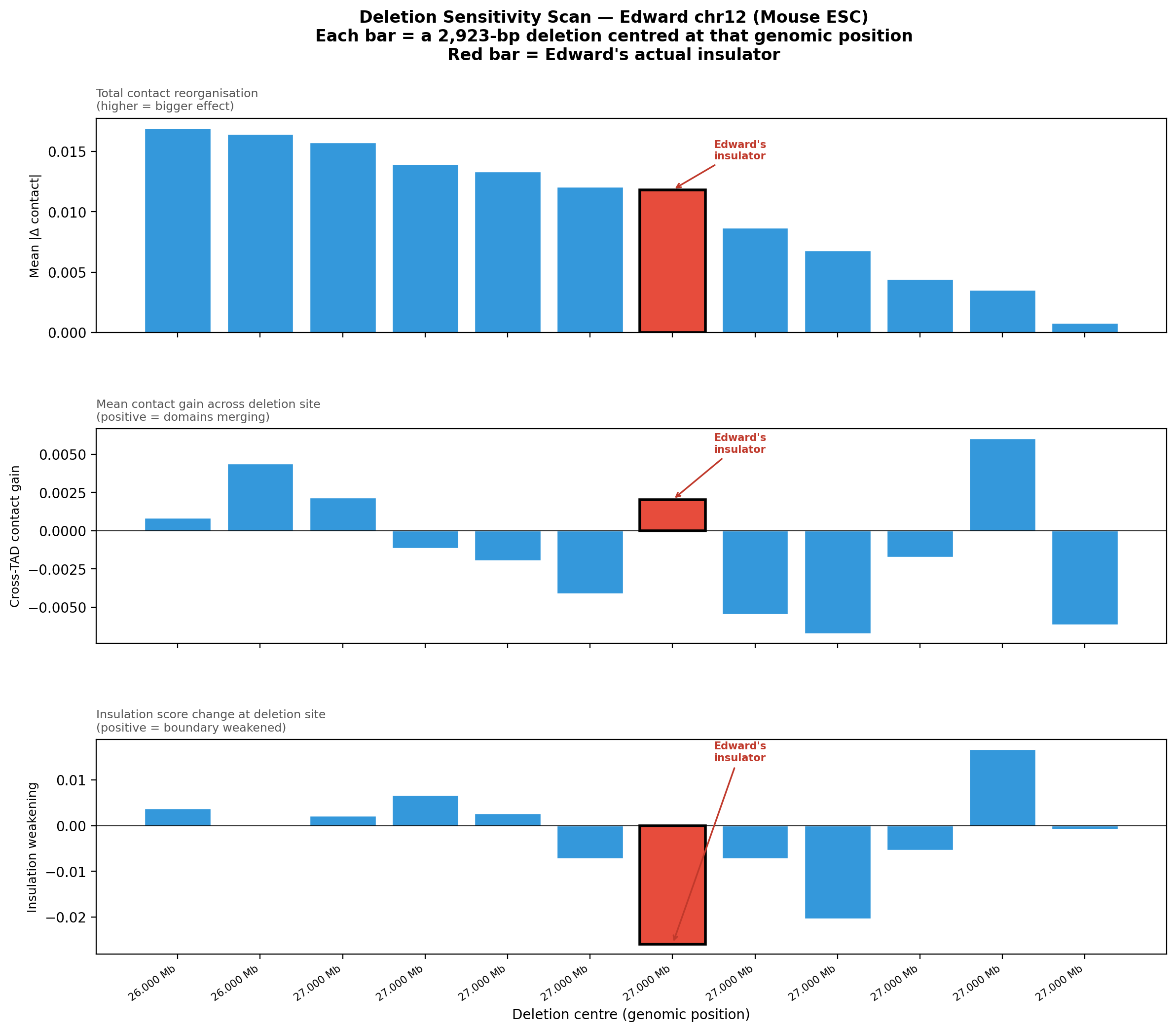
Question: Is Edward's insulator site truly special, or would any random deletion in the same region produce a similar contact-map change?
Experiment: 12 evenly-spaced deletions of the same size (~3 kb) were simulated across the 1 Mb window using AlphaGenome (Mouse ESC, EFO:0004038). The wild-type was predicted once; each deletion was compared against it using three metrics.
• Mean |Δ contact| — total contact-map reorganisation (how much anything changed).
• Cross-TAD contact gain — average gain in contacts across the deletion site; positive = domains merging.
• Insulation weakening — how much local boundary strength drops at the deletion site; positive = boundary lost.
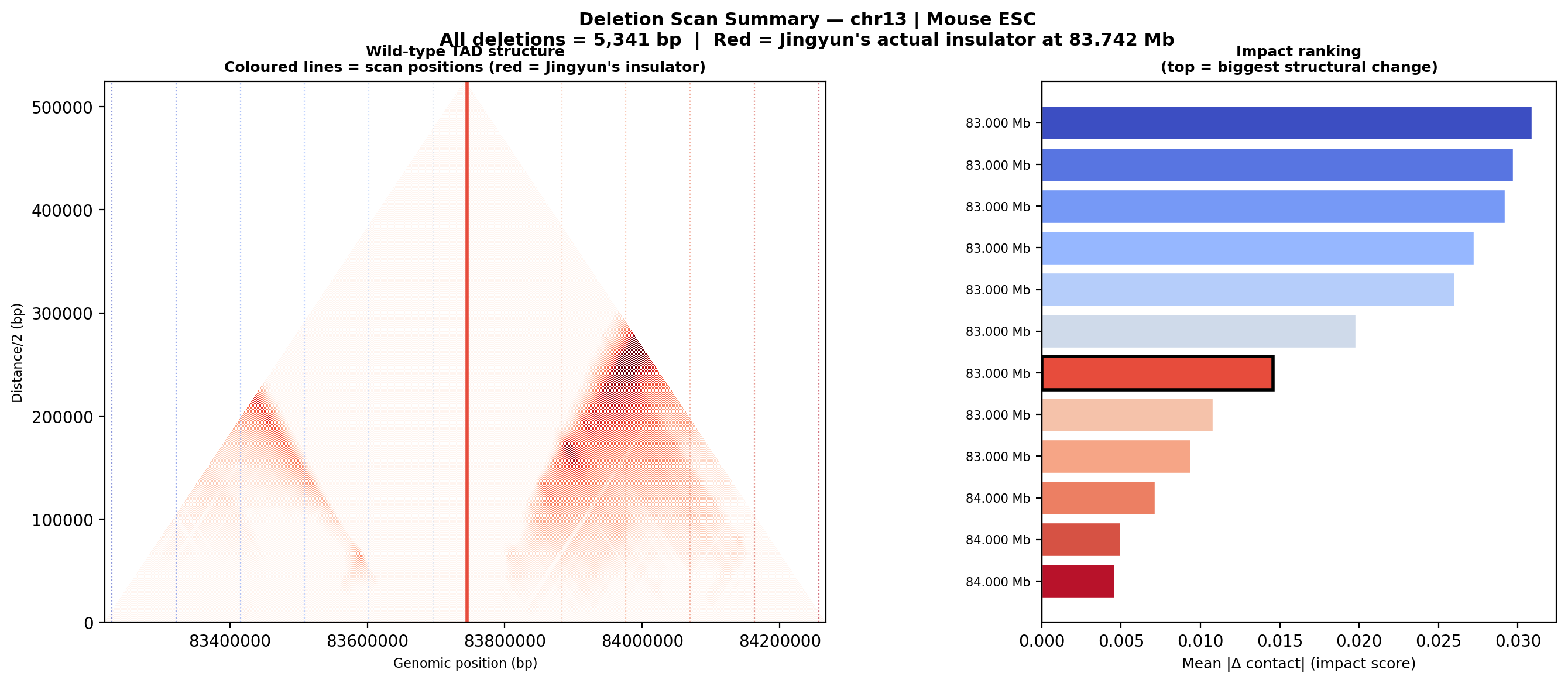
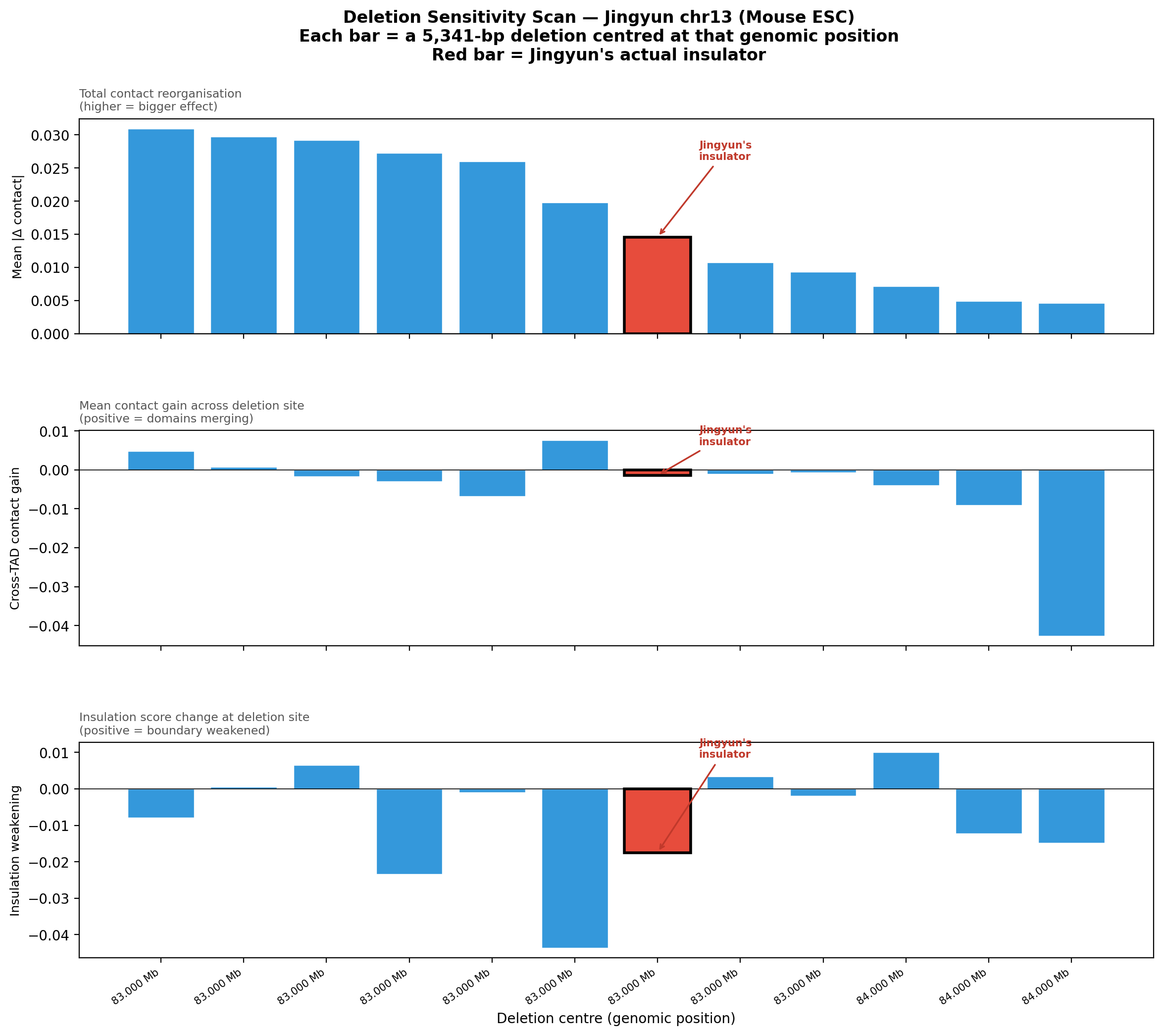
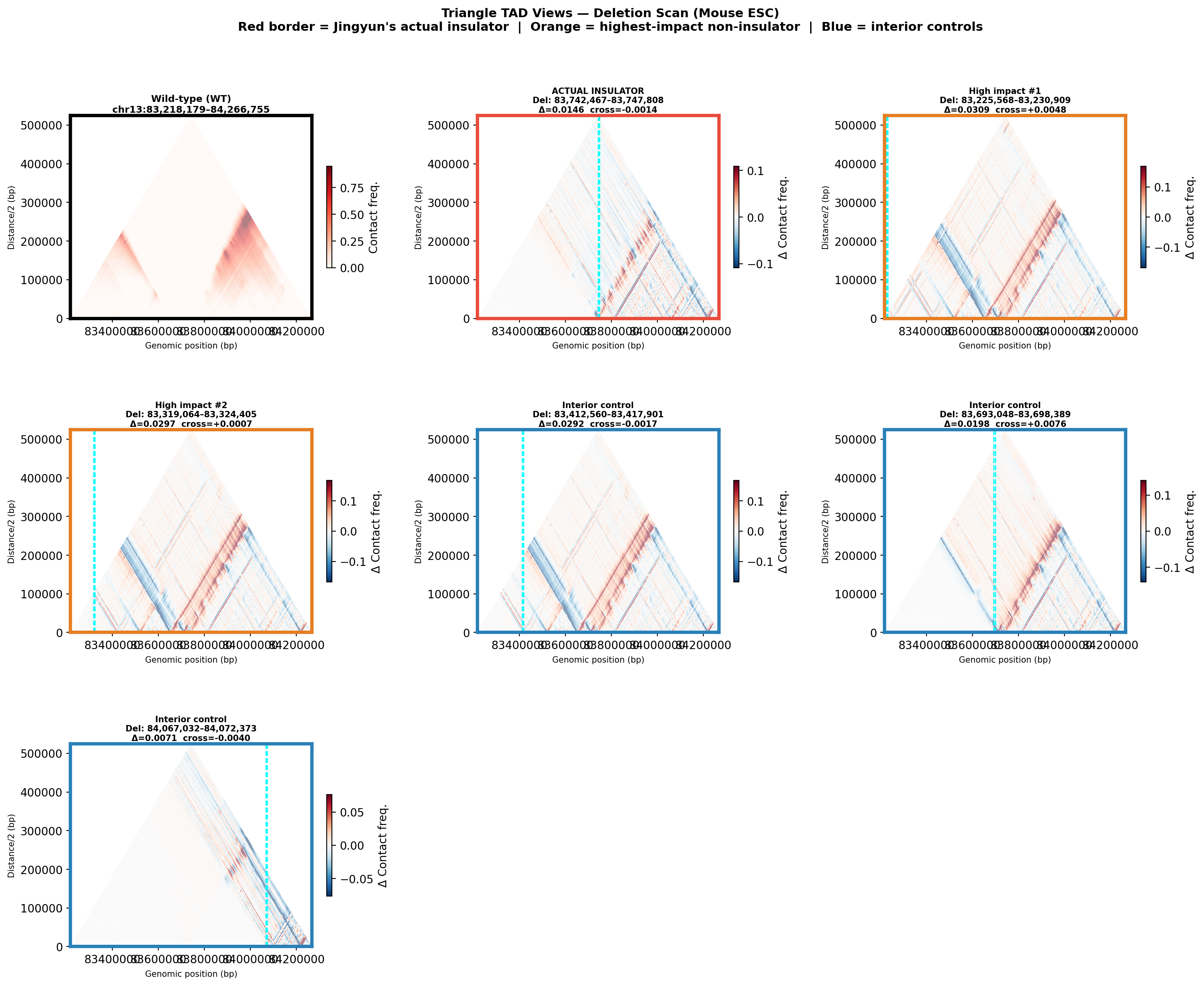
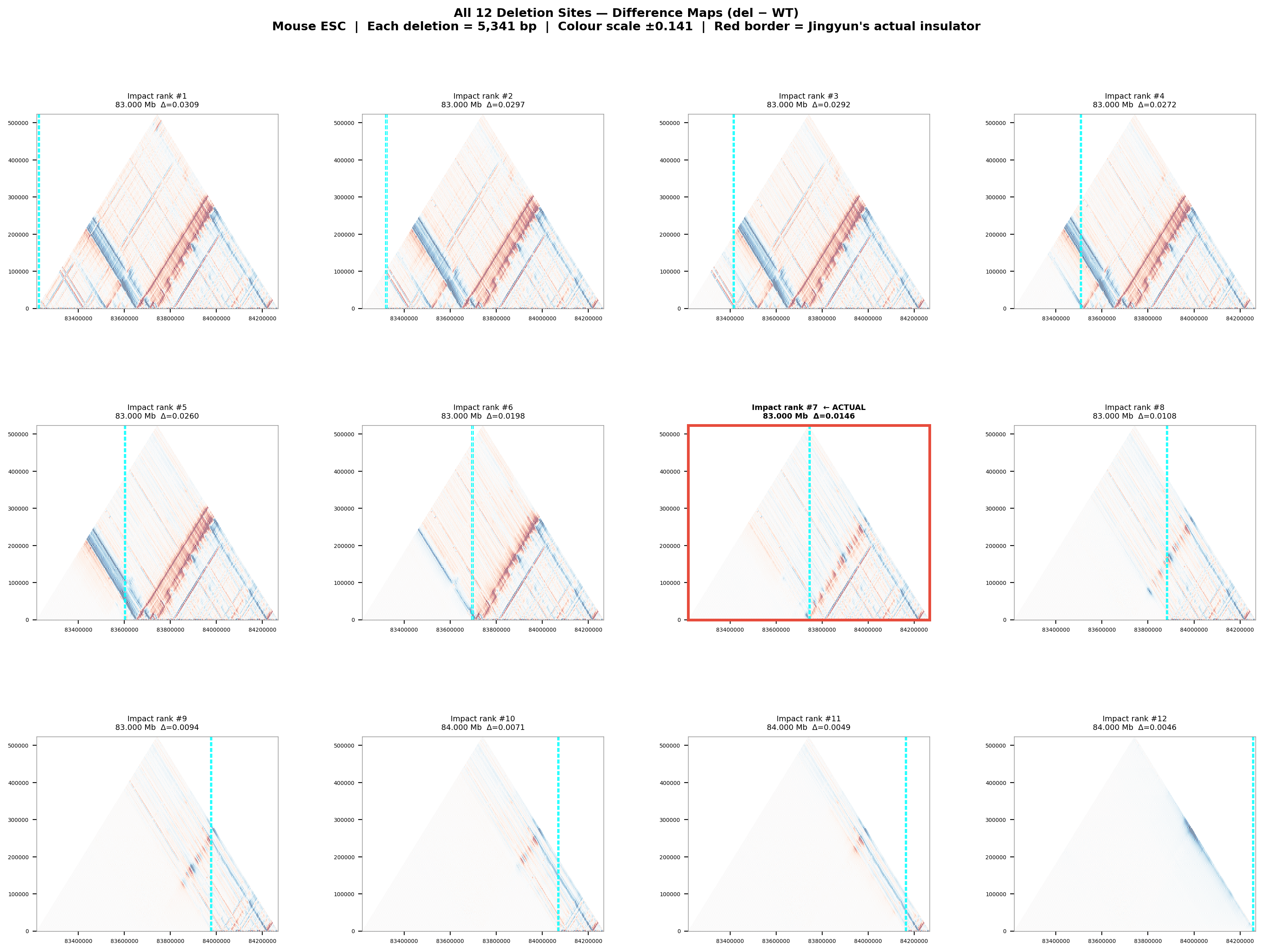
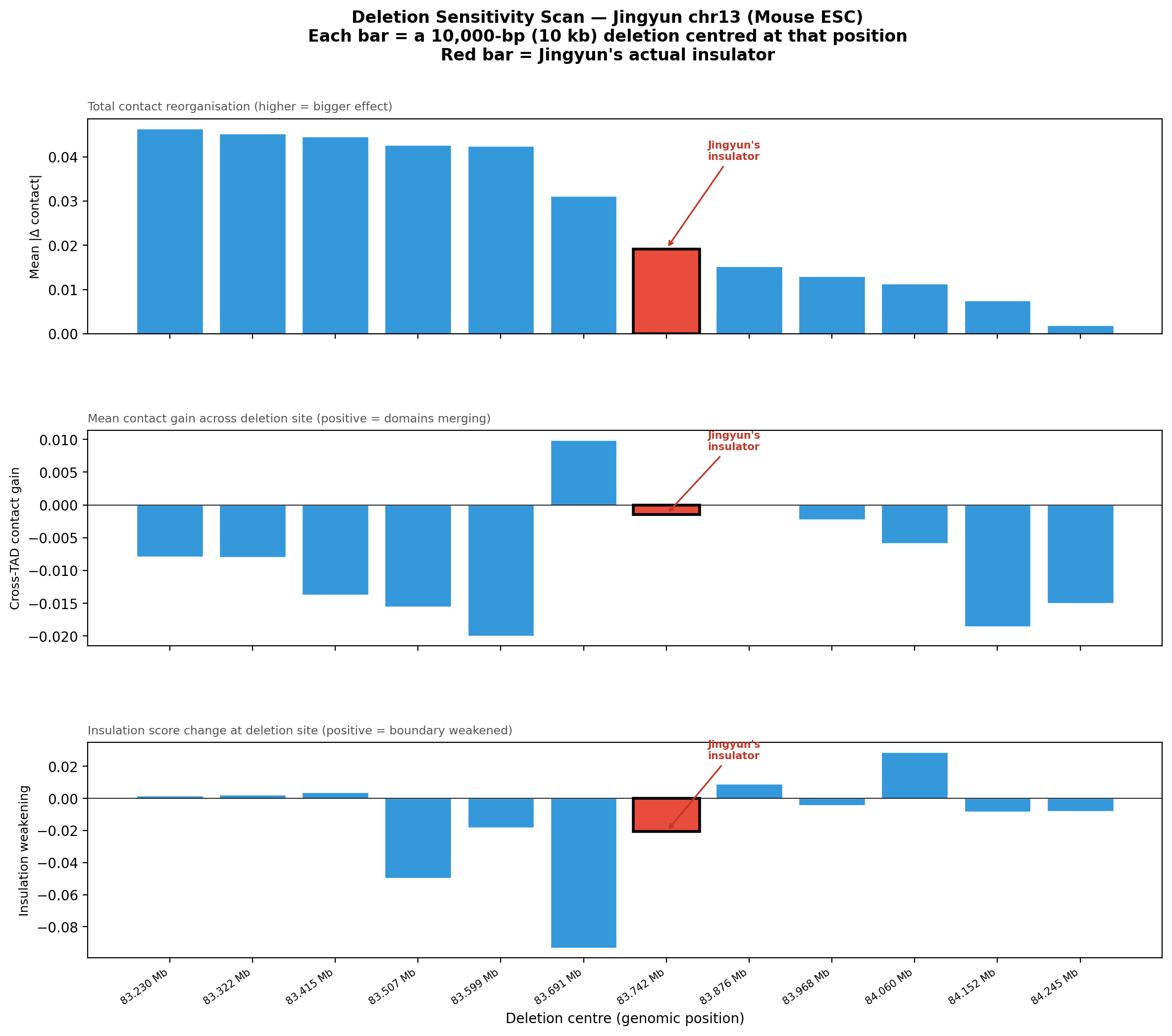
Deletion Sensitivity Scan — Jingyun chr13 NEW
Question: Is Jingyun's insulator site truly special, or would any random deletion in the same chr13 region produce a similar contact-map change?
Experiment: 12 evenly-spaced deletions of the same size (~5.3 kb) were simulated across the 1 Mb window centred on the chr13 insulator using AlphaGenome (Mouse ESC, EFO:0004038). The wild-type was predicted once; each deletion was compared against it using three metrics.
• Mean |Δ contact| — total contact-map reorganisation (how much anything changed).
• Cross-TAD contact gain — average gain in contacts across the deletion site; positive = domains merging.
• Insulation weakening — change in cross-boundary contact frequency at the deletion site; positive = boundary lost, negative = local reorganisation.
Multi-Size Deletion Scan — 10 / 40 / 80 kb NEW
Question: Does deletion size matter? We re-ran the 12-position scan with three larger deletion sizes (10, 40, 80 kb) to see whether the actual insulator site becomes more or less detectable as deletions grow larger.
• Edward chr12: With 10 kb and 40 kb deletions, his insulator jumps to rank #1/12 (vs. #7 with the ~3 kb insulator-size deletion), with insulation weakening of +0.45 — a massive, unambiguous boundary collapse. At 80 kb, it falls to #4 as adjacent CTCF anchors also become disrupted. The insulation weakening remains strongly positive at all sizes, confirming a true functional boundary.
• Jingyun chr13: Her insulator stays at rank #7/12 across all sizes, with near-zero insulation weakening at 40 and 80 kb. This is a sharp contrast: the chr13 site does not become more detectable with larger deletions, suggesting it may not be a dominant architectural element in Mouse ESC — or the boundary is maintained by redundant mechanisms not captured by deleting only one region.
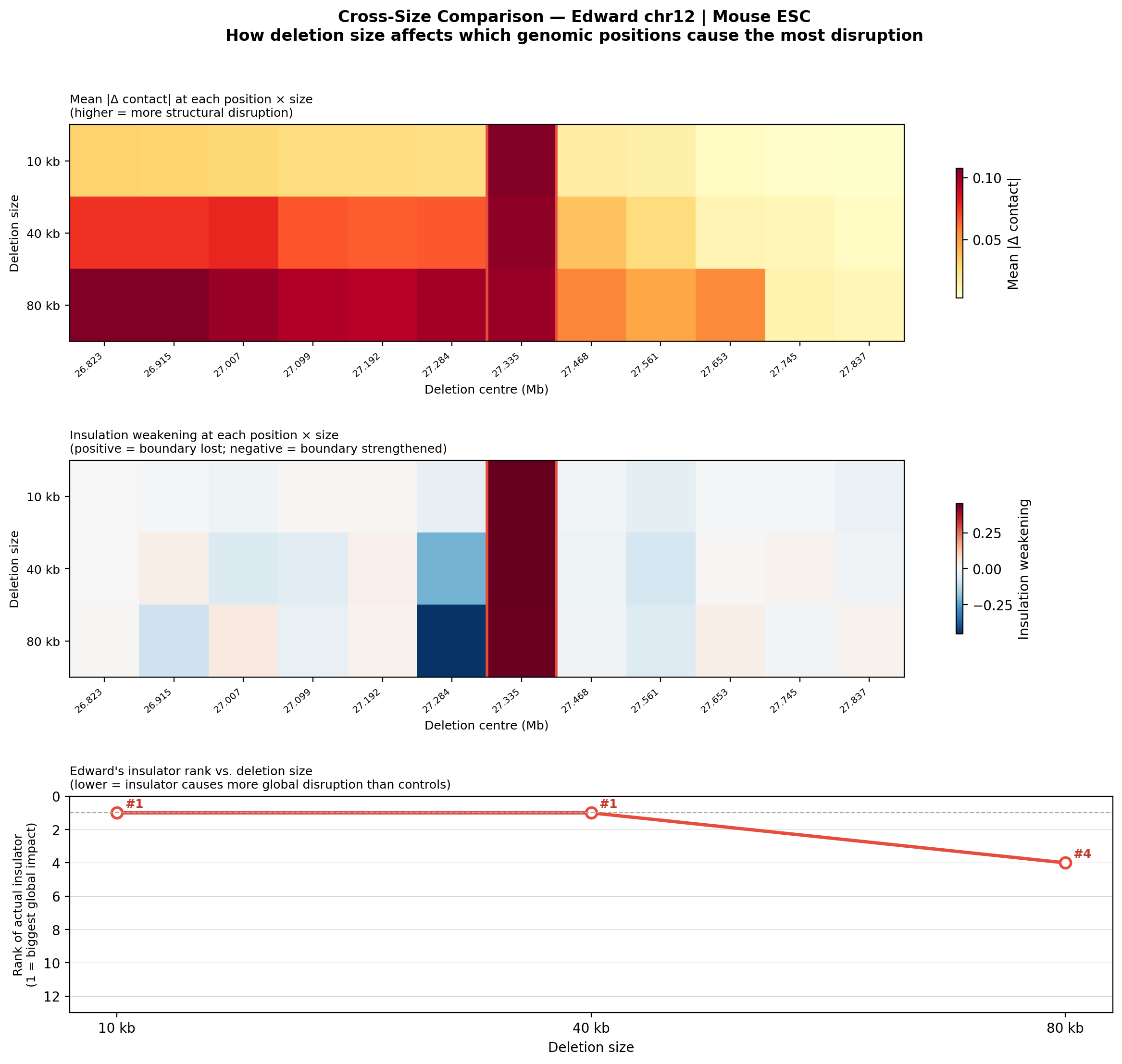
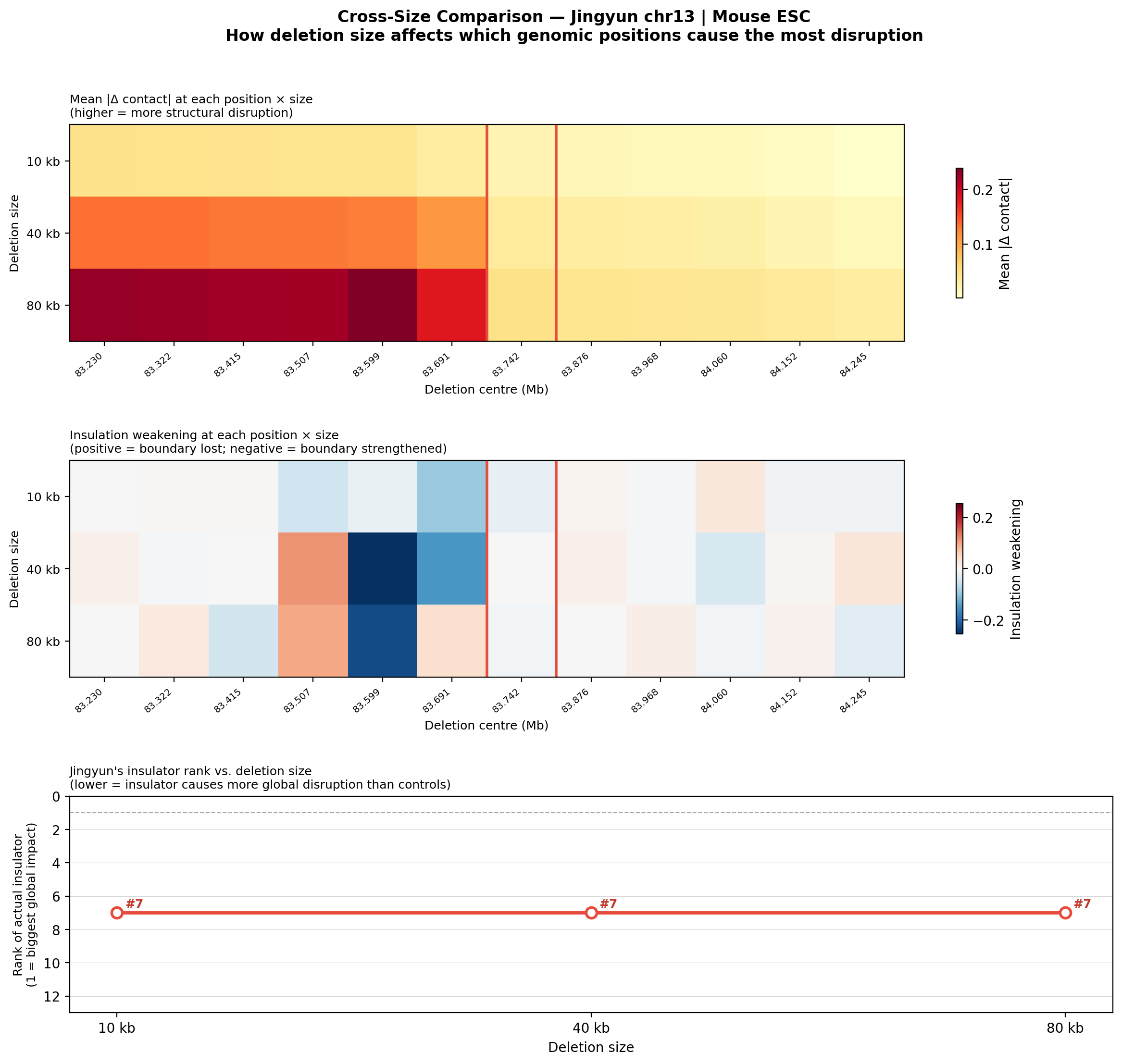
Full per-size figures (summary, gallery, montage) for all deletion sizes are available in the Edward multi-size report and Jingyun multi-size report.
Enhancer Candidate Identification & Synthetic Design NEW
The full synthetic enhancer pipeline is now running end-to-end. Starting from Micro-C contact structure, it nominates candidate bins, scores them with a dilated CNN trained on real neural ATAC-seq predictions from AlphaGenome, then generates novel synthetic sequences predicted to have high enhancer activity — using both gradient-based optimisation and motif-insertion hill-climbing.
Stage 1 — Structural nomination (Hi-C):
1. TAD membership — only bins inside a called TAD (inter-TAD regions excluded).
2. Boundary exclusion — bins within 10 kb of any strong boundary removed (insulators/CTCF).
3. Contact enrichment filter — summed contact to ±50 kb neighbours; above-median bins retained.
Stage 2 — Sequence scoring (CNN):
4. Sequence extraction — 500 bp window centred on each candidate from mm10.fa → (4 × 500) one-hot matrix.
5. Dilated CNN — three Conv1d layers (dilation 1→2→4) capture motif grammar at multiple scales; sigmoid → score [0, 1].
6. Training labels — real neural ATAC signal averaged across AlphaGenome forebrain + midbrain + hindbrain tracks (
UBERON:0001890 / 0001891 / 0002028); 224 candidates, loss 0.065→0.004, 30 epochs.Stage 3 — Synthetic design (in silico):
7. Gradient optimisation — Gumbel-softmax one-hot, Adam updates toward max CNN score; temperature annealed 1.0→0.1; GC-content penalty to maintain 45%.
8. Motif-insertion hill-climb — top real candidates as seeds; greedy insertion of Sox11/Sox2/NeuroD1/AP-1/KLF motifs over 3 rounds.
• Stage 1: complete — 161 Sox11 candidates + 76 Mir9-2 candidates called from contact structure.
• Stage 2: complete — CNN trained on real neural ATAC labels (AlphaGenome confirmed available for MUS_MUSCULUS); bars coloured green→red by score, gold ★ stars mark bins ≥ 0.5.
• Stage 3: complete — gradient designs score 0.80+ (vs 0.76 best real candidate); motif insertion adds +0.03–0.05 per round via Sox11/NeuroD1/KLF motifs.
• Next (experimental validation): AlphaGenome in-silico validation of designed sequences via sequence substitution into real locus context (
validate_designs.py), then MPRA wet-lab readout.
• Real neural ATAC labels confirmed. AlphaGenome ATAC tracks for MUS_MUSCULUS forebrain (
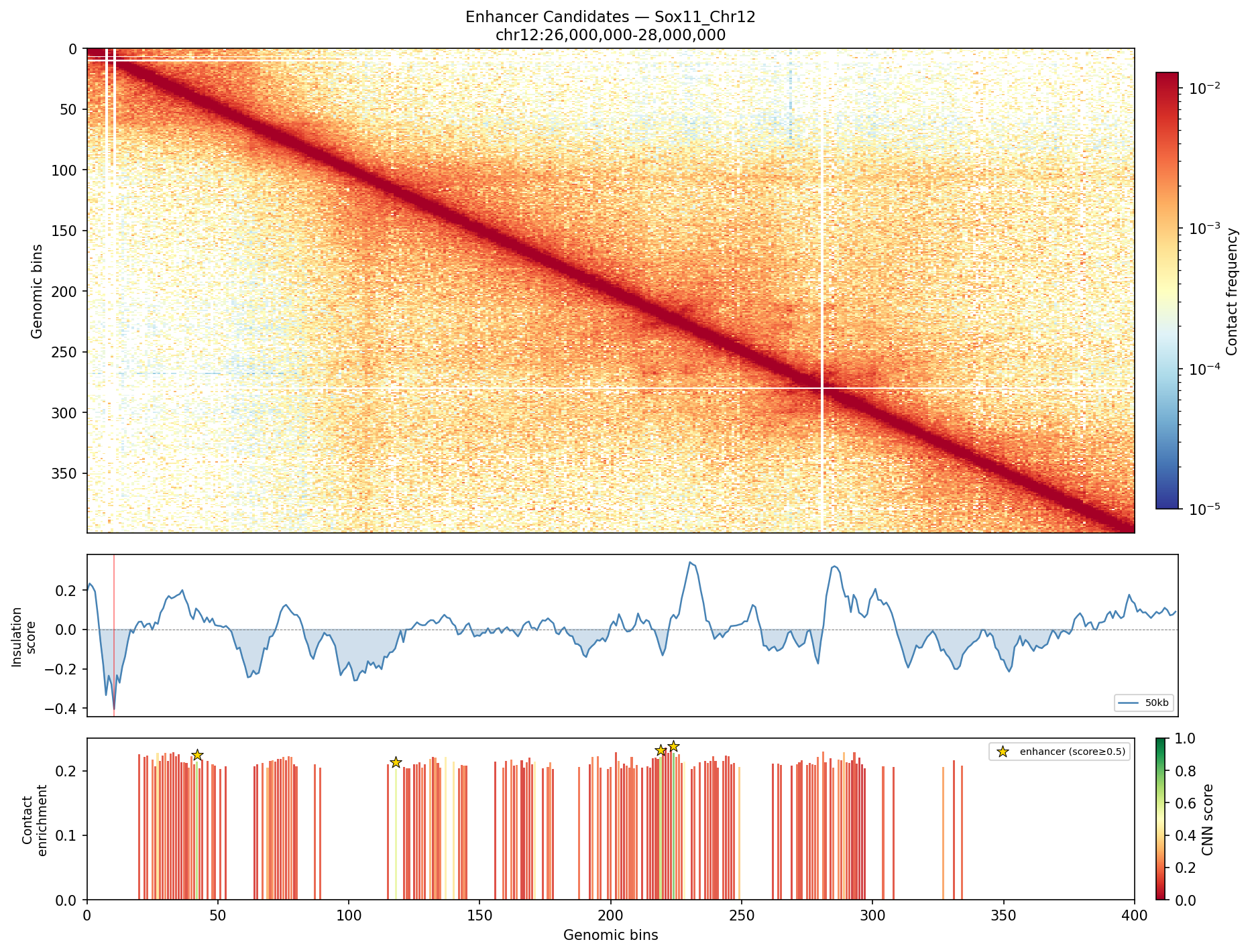
UBERON:0001890), midbrain (UBERON:0001891), and hindbrain (UBERON:0002028) are all available and returned signal. Labels averaged to a pan-neural accessibility track (mean ATAC ~0.07–0.08 across candidate bins, consistent with sparse accessibility genome-wide).• Sox11_Chr12 — 161 candidates, CNN scored. Top real candidate: chr12:27,120,000–27,125,000 at score 0.764. High-scoring bins cluster in the left TAD (~26–26.75 Mb), consistent with the dense distal regulatory landscape of the Sox11 locus.
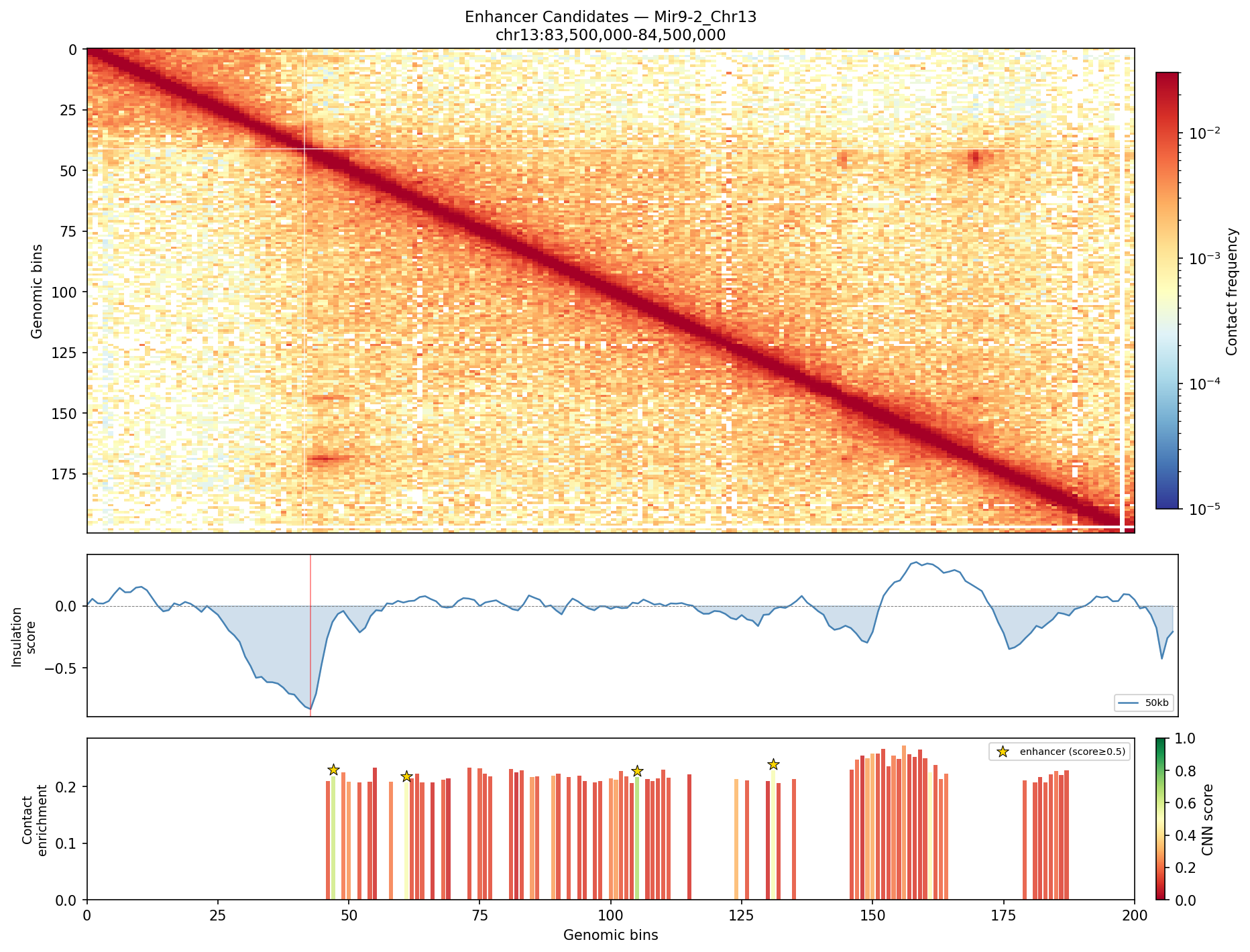
• Mir9-2_Chr13 — 76 candidates, CNN scored. Top real candidate: chr13:84,025,000–84,030,000 at score 0.677, in the right-side cluster that already showed elevated contact enrichment — a convergent structural + sequence signal.
• Gradient designs consistently outperform real candidates. All 20 gradient-optimised sequences score ≥ 0.80, above the highest real candidate (0.764). GC content maintained at 44–47% by the penalty term. These are genuinely novel sequences not present in mm10.
• Motif insertion reveals neural regulatory grammar. Each round adds +0.025–0.05 CNN score. Most accepted motifs: KLF (
CCCCGCCC), Sox2 (CTTTGTT), Sox11 (AACAAAG), NeuroD1 (CAGATGG). This identifies KLF GC-boxes and Sox-family sites as rate-limiting for predicted enhancer activity in this neural context.
Sox11 — chr12:26,000,000–28,000,000 (2 Mb · 161 candidates · CNN scored, real ATAC)
Mir9-2 — chr13:83,500,000–84,500,000 (1 Mb · 76 candidates · CNN scored, real ATAC)
Synthetic Design — Gradient Optimisation & Motif Insertion
• All 20 designs score ≥ 0.80 CNN — above the best real candidate (0.764 Sox11, 0.677 Mir9-2).
• Best design: 0.815 (Sox11 region), GC = 45.6%.
• GC content 44–47% controlled by gradient penalty; no degenerate poly-G or low-complexity outputs.
• Sequences stored in
data/processed/gradient_designs.tsv.Motif-insertion designs — 10 sequences (top-5 seeds per region, 3 rounds):
• Most accepted motifs: Sox11 (
AACAAAG), AP-1 (TGAGTCA), Sox2 (CTTTGTT), NeuroD1 (CAGATGG) — Sox11 motif accepted in every seed.• Total improvement per sequence: +0.009–+0.050 over 3 rounds (each round +0.002–+0.020).
• Results in
data/processed/motif_designs.tsv with full insertion history.Interpretation: Gradient designs identify the sequence features that jointly maximise the CNN objective — a compressed picture of what the model considers “neural-enhancer-like.” Motif insertion reveals the rate-limiting regulatory grammar for each real candidate locus. Together they give two independent views of the same underlying biology.
The papers Kelvin shared (Cell Systems 2025 + MolecularPost overview) use a two-stage CNN approach: (1) sequence → accessibility, then (2) transfer-learn to in vivo enhancer activity. The full pipeline is now implemented and running: candidates identified, CNN-scored with real neural ATAC labels, and synthetic sequences designed by gradient optimisation. The designed sequences consistently score above any real candidate, meaning the model has found a region of sequence space predicted to be more active than the endogenous regulatory elements it was trained on.
The next step is experimental validation: MPRA (synthesise ~200 sequences including gradient designs, motif-insertion refinements, and real candidates; transfect into neural cells; measure reporter output by sequencing) or AlphaGenome in-silico validation (substitute each designed 500 bp into the real locus context and compare predicted ATAC delta). Once real ATAC-seq arrives from Jingyun & Ed's Dovetail experiment, retraining will sharpen CNN specificity and the design loop will restart with higher-quality labels.